To date, there has never been a confirmed case of second cousins (2C) or closer that don’t share DNA. There have been a few rumblings here and there, but nothing proven. See “Second Cousins (Or Closer) That Don’t Share DNA?” for more details.
But what about second cousins once removed (2C1R)? That’s only a single meiosis away from a 2C relationship. Is it possible to not share DNA at that distance? If I don’t share DNA with my 2C1R, should I suspect a misattributed parentage event, or is that normal? Is there anything I can do to give myself some peace of mind?
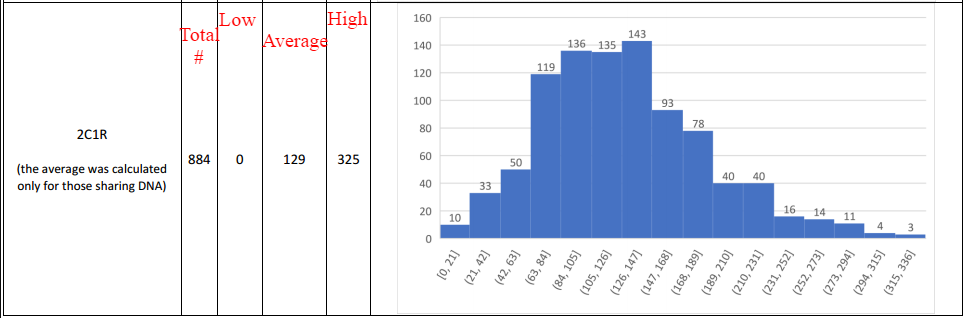
According to data from the 2016 update to the Shared cM Project, several submissions to the project reported no shared DNA between 2C1R. The histogram shows the distribution, and how often 0 cM shared is for that relationship:
Sharing no DNA at a distance of 2C1R appears to be possible, assuming the submissions were correct. But is that the end of the analysis?
A Real-Life Example
I was recently corresponding with someone on a mailing list who had a 2C1R relationship that doesn’t share DNA. She had tested a handful of other close relatives who all matches that 2C1R, but she didn’t.
It was a perfect case to examine this relatively rare situation, and she was kind enough to let us examine it on the blog for everyone to learn! Thank you!
The Family
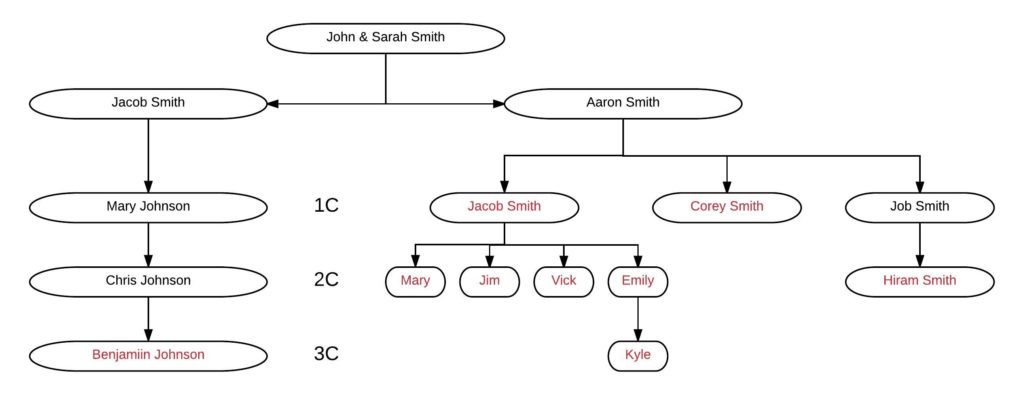
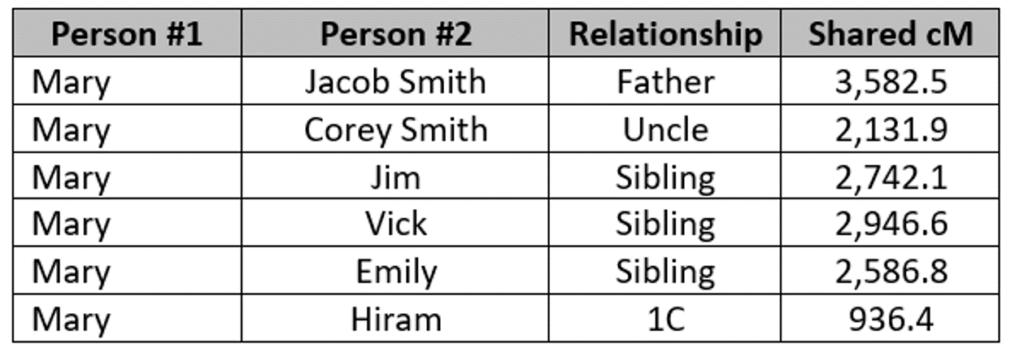
Mary [all names are changed!] has tested herself, her 2C1R, and six other members of the family that we’ll look at in this blog post. The family tree looks like this, where the people in red are tested, and their segment data is available:
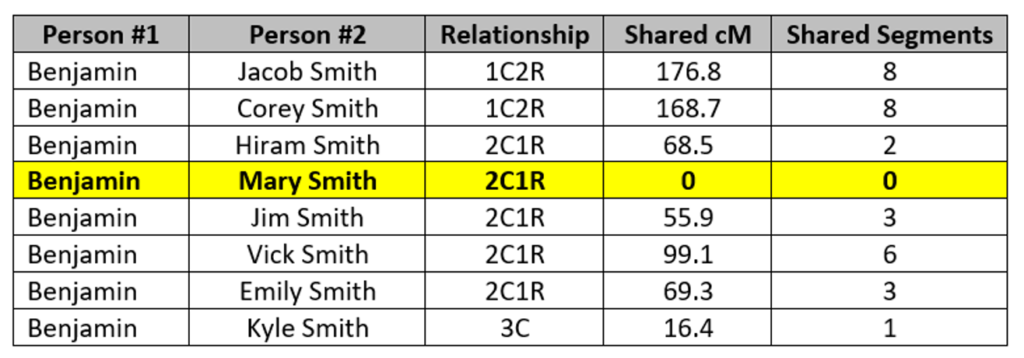
The key relationship here is Mary Smith and Benjamin Johnson, who are 2C1R. They share no DNA in common, which as noted above is relatively rare but not impossible. Every other tested person in the tree shares DNA with Benjamin, ranging from a low of 16 cM to a high of 177 cM:
Mary’s three siblings, for example, share between 56 and 99 cM with Benjamin, but Mary shares no DNA with him!
Mary’s Relationship to Her Father and Siblings
The very first thing to check is Mary’s relationship to her immediate family (father, siblings, uncle, and 1C). If there is a misattributed parentage event that explains Mary’s lack of sharing, this comparison will make it clear. However, the comparison shows that Mary is related to her immediate family as expected:
Clearly, there are no problems with any of these relationships. The shared DNA with the uncle is on the high side, but not outside the range of the Shared cM Project (range 1301 – 2193 cM, average 1744 cM).
Accordingly, just this analysis alone suggests that Mary’s lack of sharing with Benjamin is pure chance.
To the Segments!
With the segment data, we can examine this phenomenon in greater detail.
For example, let’s test this situation using Mary and her three siblings. We know that the siblings all share between 3 and 6 segments (56 and 99 cM) with Benjamin, the 2C1R. And we know that Mary shares no DNA with Benjamin. So what can we hypothesize with regard to Mary and the segments her siblings share with Benjamin?
We can hypothesize that Mary should not share any of those segments with her siblings. If she did, we would expect Mary to match Benjamin on these segments.
This will be a little difficult to test with siblings since we can’t tell which segments came from which parent, and we’re only looking at the paternal side of the family. But we can hypothesize two things:
- Mary and her siblings cannot share fully-identical regions (the green regions using the GEDmatch One-to-One tool) at any location where her sibling matches Benjamin; and
- If Mary and a sibling share a half-identical region (the yellow regions using the GEDmatch One-to-One tool) at any location where her sibling matches Benjamin, that segment must be a maternal segment!
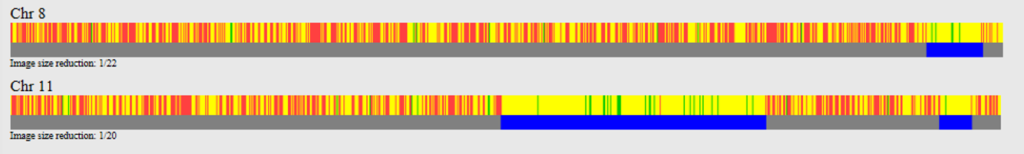
So let’s do a comparison of Jim and Benjamin, and then Jim and Mary. Jim and Benjamin share DNA on chromosomes 8 and 11:
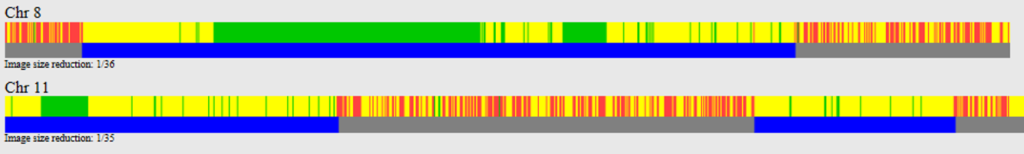
As expected, there are no HIRs shared between Jim and Mary at the locations where Jim and Benjamin share DNA. On chromosome 8 it is clean and simple; Mary and Jim share no DNA where Jim and Benjamin share DNA.
On chromosome 11, there is about 10 Mb of overlap between the large segment shared by Benjamin and Jim (top image) and the segment shared by Jim and Mary (bottom image). This region is likely a maternal match between Jim and Mary.
Second Cousins or Closer?
The process above is what would be needed to prove that there are 2C or closer that don’t share DNA. It may happen eventually, if millions of second cousins are tested, and hopefully this will provide some insight into how to analyze a suspected case.
Since I’m just getting started with DNA matching, this is a good reminder that sometimes, just sometimes, exceptions apply! Thanks and I’m enjoying learning from your posts.
Thanks for another interesting analysis.
It would be helpful to include labels on the first chart so that we know what the numbers represent.
Also, a bit of clarification re hypothesis 2 – … that segment must be a maternal segment!
Can you elaborate on which side of the family the maternal segment match may come from.
Thanks
Love the article. Great work as always, Blaine.
If you have Mary and her siblings and their Dad, can’t you phase their DNA to separate their maternal and paternal matches?
If they do not share dna when they should, then look to see if there is an X chromo (23) sharing?
Fascinating. Thank you for the insight. Were all of them tested with the same version chip? That could make a small difference.
Still learning this stuff. I’d find it helpful to include arrows or clearly label the sections of DNA being described in the analysis.
Is this because when a male and female DNA combine to create the new organism, the snippets of DNA taken are in chunks, e.g., 10 contiguous sets from a particular chromosome are.
I share 28 CM with my 2c1r – all my second cousins (3 in no) share over 100-355. Not an example of no DNA but certainly an example of the randomness of selection.
My sister and I do not match a paper trail third cousin (who is actually a one and half times third cousin due to two half-sisters marrying two brothers) and neither does our aunt in that line who would be a 2C1R to our third cousin. However, my sister and I do have some common shared matches with her up to about 20 cM. Our connection is in Ireland not in a colonial community. Can we use this shared match information to validate our relationship?
Blaine has shown what it looks look when people in a 2C1R relationship don’t share DNA–and what it takes to prove that random chance is the cause rather than a misattributed parentage or a mistake by the genealogist. But what are the overall odds of it happening? ISOGG has data (old but presumably reliable) to puts the odds at about 1/1000. Blaine’s user-submitted data suggests it might be slightly more common than that.* It’s rare, to be sure, but something we should expect to see every once in a while. http://isogg.org/wiki/Cousin_statistics#Theoretical_probabilities
It’s also worth noting that Blaine’s data includes half 2nd cousins, which is similar to 2C1R in that it is, on average, a 2nd cousin relationship divided by two. (The difference lies in where the extra division takes place, either structurally in the first generation–a single shared great-grandparent with two partners rather than two great-grandparents in common–or in the last generation–a 2nd cousin relationship with an extra meiosis/generation down one of the two lines, therefore removing on average half of whatever DNA had overlapped at the 2C level). Notably, for half 2nd cousins the minimum in Blaine’s data set is also 0 cM.
The fundamental question ultimately being asked by inquiries like this is, what is the minimum number of divisions (structural or meiotic) after which two genealogically related individuals might no longer be related genetically. Thanks in part to Blaine’s research, we’re zeroing in on an answer between 5 and 7, depending on the particular relationship pattern in question. For example, ISOGG’s data shows that there is a .01% chance a person shares no detectable DNA with a 3x-great-grandparent. That is 5 meiotic divisions down a single line. 2nd cousins represent six divisions, three down each of two separate lines. 2C1R and half-2C are both seven divisions. (Incidentally, looking through all the data on the ISOGG page, it appears a median might be around 11 divisions, equivalent to a 4C1R relationship. That is, a person could expect to be genetically related to about half of his/her genealogical cousins at this distance. The other half would share part of a genealogical pedigree but no longer share any IBD DNA segments.)
*Blaine’s data for 2C1R lumps people who share 0cM with others who share as much as 21cM, so it’s unclear exactly how many of the 10 relationships in that “bin” shared exactly 0cM. Perhaps it’s just one or two of the ten cases, which wouldn’t be far from the odds posted by ISOGG (that is, 1/884 or 1/442). Blaine presumably has the full data and could tell us.
Very interesting. I’ve just discovered this situation in my own family. My second cousin and I have both tested at Ancestry. We only share 56 CM across 6 DNA segments, which seems low for a second cousin relationship. My daughter and son have both tested at Ancestry, also. My daughter is not a match with my second cousin, but my son is a match.
On the “flip side” of this question: I am interested in everyone’s thoughts on the impact of “double cousins” within a family. Should I assume that a double cousin relationship will show twice the predicted shared DNA range for first cousins? Is this the most likely cause of a distant cousin (5th cousin through confirmed trees) with shared cM that is outside the range predicted in the Shared cM project chart? My 5th cousin and I test at 83 on 2 segments (Ancestry), which Shared cM suggests is closer to 4th or even 3rd cousins. This is in a Norwegian family (in the US since 1890) and no known double cousins in my side, but possibly on the cousin’s line. Is this the best place to start investigating?
Nice!
I have this same situation seeking my bio paternal grandfather.
Testing one and she matches most all my dna matches — but at higher cM than me.
I suspect now we are 2C1R..
We had thought we were 1/2 first cousins.
My dad was an NPE event .
We suspected her grandafAther as my bio-grandfather .
She is now going to try and download her ancestry dan then upload dna to gedmatch…
OOPs sorry she shows NO dna at all with me.
Does not even show up under common matches.