This is the first entry in a two-part series describing my recent experiences with genetic testing through 23andMe. Although I was most excited by the genetic genealogy information provided by the results, I thought that readers might be interested in some of the health and trait information as well. If I forget to discuss something, feel free to ask in the comments and I’ll do my best to respond.
Note that this discussion is limited to a cursory analysis of my results. Anyone who is considering testing through 23andMe should be aware that scientists are only just beginning to study and understand the connection between genetics and health, and thus the results are not meant to be interpreted as absolutes. Be sure to analyze your own results with this caveat in mind, and do your own research into the testing process.
23andMe
Since many others have discussed the 23andMe kit and the spitting process, I won’t go over that here other than to provide a picture of my actual testing kit:
Although the typical processing time is 4 to 6 weeks, my sample took a little longer to process because 23andMe was still working with the New York Department of Health regarding samples collected in New York. Lucky for me I was very familiar with this situation and had decided to collect my sample outside New York.
Health and Traits
In addition to ancestry, 23andMe’s SNP results are examined for information about over 90 different types of health and traits. I found the health and traits analysis to be very interesting. Some of it, based on my own knowledge of my family history, was unsurprising. As important as genetics may be in the future for predicting predilection for disease, there will always be a place for family health history (and apparently the U.S. Surgeon General agrees). There were, however, some health and trait results that I did not expect, especially regarding gene variants that can influence genetic disorders in females (and thus might be something to keep in mind if I have a daughter).
Blood Type
Interestingly, 23andMe does not report on ABO blood groups even though the v2 chip collects this data (although 23andMe is planning to do so in the future). For example, I knew I was Blood Type A, but I didn’t know if I was AA or AOÂ A member of the 23andMe community – which I’ve spent a great deal of time perusing – provided directions for deciphering blood type. Turns out I’m AO.
It was also fun to confirm the genetic basis behind some of my physical characteristics. I have wet earwax, if you were just dying to know. You can read more about the earwax gene at the NYT and at Gene Expression. I can also taste bitter, meaning that perhaps I shouldn’t like raw broccoli and brussels sprouts as much as I do!
CCR5
Some time ago I wrote “Are you immune to HIV and smallpox?” about the CCR5 gene. A certain variant of the CCR5 gene has a 32 base-pair deletion (CCR5-delta32) and has been suggested to provide a measurable degree of protection against certain viruses such as the HIV virus. Scientists theorize that it might have been selected for during viral plagues in human history. The CCR5-delta32 allele is nearly absent among Amerindians and East Asians but is found in some African populations. The allele is found in high concentration among Eurasians. Indeed, the average frequency of the allele among European populations is estimated to be as high as 10%.
Since my ancestry is largely European, I thought it was possible that I had at least one copy of the CCR5-delta32 allele. However, my 23andMe results show that I have two intact versions of CCR5:
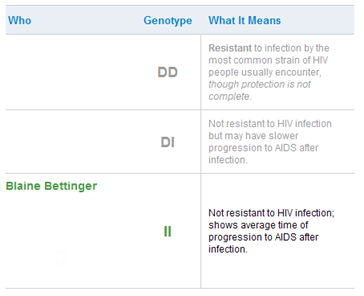
Type 2 Diabetes
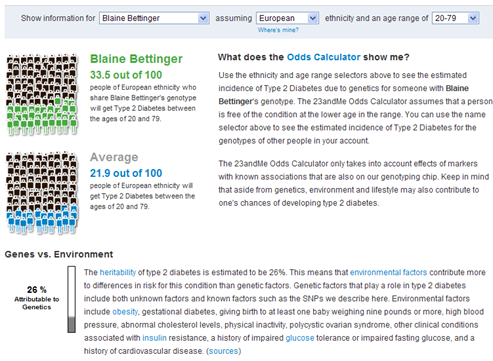
The 23andMe test also examines 9 SNPs believed to be related to Type 2 Diabetes. As shown below, my results suggest that I might be more prone to Type 2 diabetes than the average European; my estimated incidence is 33.5 out of 100 compared to 21.9 out of 100. There is diabetes in my family, so this result was not unexpected. Current understanding is that genetic factors account for only 26% of the risk of diabetes.
So what do these results really mean for me? Exactly the same thing that a family history of diabetes means – I need to eat healthy and exercise (and so do my children).
You might ask yourself why I feel comfortable posting this information online, specifically where an insurance company could find it. The answer is two-fold; first, for a number of reasons, I believe that the fear of insurance companies using our genetic data against us is overblown; and second, the insurance companies already obtain this information in an slightly more abstract form when they ask about family history. With the Genetic Information Nondiscrimination Act, however, health insurers will no longer be able to ask about family history, precisely because family history belies genetic information (note that life insurers and other types of insurers WILL still be able to inquire about family history and genetic tests).
Browse Raw Data
Another great feature is the ability to browse your raw SNP data, including the results that have not yet been associated with any disease or trait (which is the vast majority of the SNPs). The data is broken into 22 pairs of chromosomes, the sex chromosomes (XX or XY), and mtDNA. The data can be searched by gene or SNP name, or can be browsed randomly. This feature is useful whenever I come across a new study that suggests a link between SNP(s) and a genetic trait or propensity for a genetic disorder; I can look up my own result for this particular SNP.
I can also download all my own data. I used this option to download my Y chromosome SNPs for Y-Chromosome Genome Comparison, which I’ll discuss more of in a later post.
Conclusion
This post included just a few thoughts about my testing experience, excluding my ancestry results. Tomorrow’s post will examine that aspect of the testing.

How do you download the data? We also tested with 23andMe.
Bob – if you go to “browse raw data” under “ME” on the “welcome to you” page, you’ll go the new page and you’ll see “download raw data” on the upper-righthand corner. Just click on that and follow the directions from there.
Blaine –
Did you purchase the test yourself or were you provided one free of charge by 23andMe? If the latter, then it would be helpful to disclose that as part of your review.
I requested the test from 23andMe so that I could conduct an independent review of the product for my readers, many of whom have expressed interest in it. They were nice enough to provide a test without cost for my review. I’ve also assisted a number of people in analysis of their own results.
I am very impressed by this test. I also was surprised by the differences in your percentage results as posted by ancestrybydna which I tested with. This will definitely be my Next project.
Hope all is well Blaine.
Shoshone
Leery of the odds accuracy of 23andMe, I am curious as to the chance of my spit sample being mis-labeled with some other sample and results being of dire consequences. Can you ease my mind?
Avis – I cannot, of course, make any sort of guarantee about proper labeling other than to say that I know that 23andMe has safety mechanisms to prevent this type of situation. Regarding the accuracy of results, I know that several people have tested through more than one company and found that their results were identical or nearly identical (i.e. just a handful of differences among 500,000 SNPs).
Is there anyway to buy or obtain this test or one like it for myself?
hey thanks for sharing!
I tried the 23andme service and video-blogged my experience. http://bit.ly/aKereQ
23andMe reported that I, a White woman of European descent, had… 0% European. The health report seems randomly generated. The traits report gives bizarre results. There are many haplogroups. After only taking samples from four population groups, they assign you to one. When I asked on the forums what happened, a bunch of bullies seemed to try to drive me to suicide. I want my money back.