 Imagine the following scenario:
Imagine the following scenario:
- You’ve just received an email that your DNA test results are ready, and you log into your account. The welcome screen guides you through a tutorial and presents you with several tabs to choose from.
- You click the first tab which reads “Your Ancestors.” The page shares information about 35 of your ancestors from the past 300 years, identified because you have inherited some of their DNA, although you have not yet provided any genealogical information to the testing company. Each of these ancestors has their own profile page complete with dates, family members, and other information such as computer-generated images and a health report which are based on a genome reconstructed entirely from modern-day descendants.
- You then click on the tab that reads “Your Reverse Family Tree,” which contains a partial family tree that has been constructed by the testing company. Based on extensive and well-documented genealogies, there is likely only one way in which the 35 identified ancestors can fit together in a tree (although other possible combinations are provided along with statistical probabilities). There are a considerable gaps, especially on your recent immigrant grandmother’s line, but the tree appears to be entirely consistent with your many years of traditional genealogical research. Well, except for the family of John G. Rogers from the 1850’s; you’d copied that off the Internet years ago and never confirmed for yourself anyway.
- Next you click on “Your Cousins,” which contains numerous close and distant relatives in the database. Some of these cousins are Genetic Cousins (with whom you share DNA), and some of whom are Genealogical Cousins (with whom you share a genealogical relationship based on your generated family tree). There are numerous 2nd and 3rd cousins matches. There are also pending offers to join several citizen science and family research groups, including the “Descendants of Calvin Lane of Old Lyme, Connecticut” group, the “Family of German Immigrant Johann Kehl” group and the “Relatives of the American Franklin Family” group, each of which has a slightly different research goal.
- Lastly, you click on “Your Memberships,” which offers – among other things – a discount membership to the Daughters of the American Revolution based on your predicted descendancy from Revolutionary War veteran Jedidiah Johnson (although you don’t happen to share any of Jedidiah Johnson’s DNA, he’s in your generated family tree with an extremely high probability (95%)).
While the scenario I described above may sound like science fiction, it’s the inevitable future of genetic genealogy and is much, much closer than you might think (okay, maybe not the DAR offer!).
Next month at the American Society of Human Genetics 2013 meeting, researchers from AncestryDNA will present their work detailing the reconstruction of portions of the genomes of an 18th-century couple using detailed genealogical information and Identity-by-Descent (“IBD”) DNA segments from several hundred descendants of the couple in the AncestryDNA database. In other words, researchers identified several hundred descendants of a certain couple living in the 1700s and then used the DNA shared by those descendants to recreate as much of the couples’ genomes as possible.
The abstract is entitled “Reconstruction of Ancestral Human Genomes from Genome-Wide DNA Matches,” and is freely available to the public. Unfortunately the abstract does not identify the 18th century couple in question. Hat-tip to Kathy Johnston for sharing this abstract on Facebook!
How Does it Work?
Imagine John and Jane Smith, living in New England in the mid-1700’s. They have twelve children, ten of whom lived to adulthood. They now have many, many thousands of descendants living today, a handful of whom possess random segments of DNA handed down from John and Jane Smith. As the researchers note in the abstract, “[i]f such an individual had many children, leading to a large number of descendants today, much of the ancestral genome will be present in modern populations.”
These segments of DNA are identified using well-researched family trees, and are then woven together to create as much of the couple’s genome as possible. Some segments will be lost forever, although sometimes these missing segments can be derived or estimated.
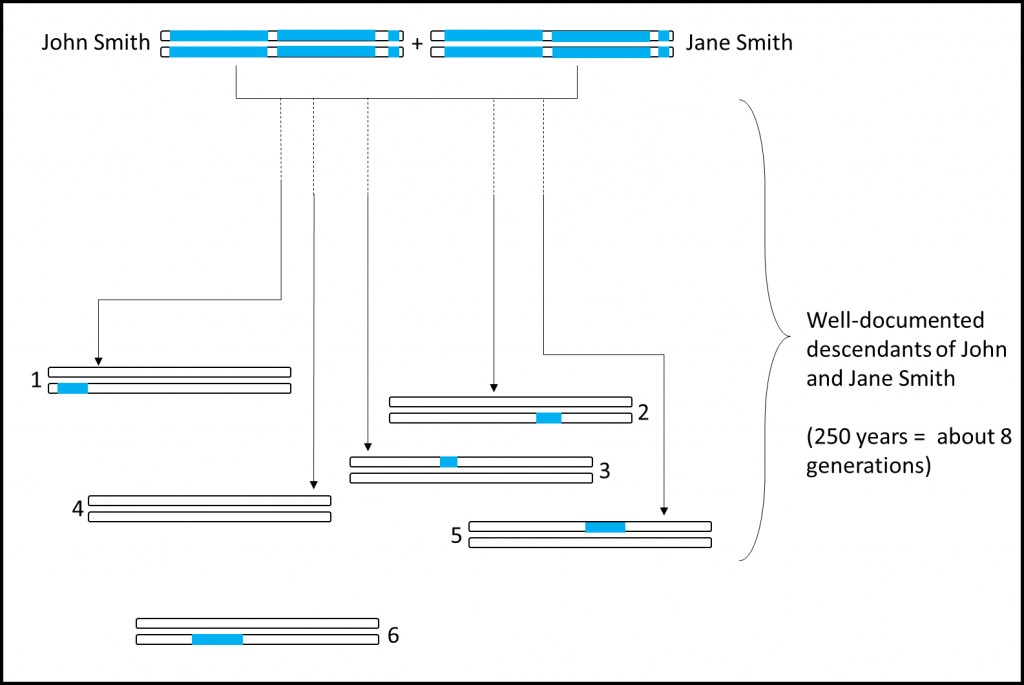
Segments of DNA from a couple in the mid-1700s are found in living descendants. Some descendants (#4) will have none of their DNA. Other relatives (#6) may have no documentation that they are related to the couple.
Slowly, the genomes of hundreds and possibly even thousands of early ancestors will be generated as millions of DNA samples and family trees are entered into a massive database. While there will certainly be numerous errors resulting from both faulty trees (either poor quality trees or trees that are wrong because of non-paternal events), and from improperly assigning shared segments to one ancestor versus another (since so many of us share many different ancestors), most of these errors will be resolved over time as more samples and trees are entered into the system and processed, and as genealogists and citizen scientists pour over the trees.
Interestingly, sometimes these recreated genomes will belong to unknown or unidentified ancestors. For example, there may be shared segments of DNA that appear to be from a couple that probably lived in the area of Boston and had children in the early 1700’s, but there is no known couple in existing records.
To be successful, this future will of course require numerous, wide-ranging, and extremely well-researched family trees as well as DNA samples from millions of individuals.
But How Can That Be Used To Generate Family Trees?
Once the genomes of hundreds or thousands of 17th, 18th, and 19th century ancestors are created and collated into a massive family tree, they can be used to recreate the genealogy of a test-taker.
For example, based on your descent from the 35 ancestors identified only by your DNA, there may be only a single way to collate those ancestors into a single family tree; only one line of descent from all of these 35 different ancestors. Alternatively, there may be multiple different possible trees that are presented to the user with a probability. The service may even suggest having a certain relative tested (we suggest having a descendant of your great-grandfather – your second cousin – tested to further refine this family tree), or it may ask you a series of questions to more accurately resolve the conflicts in the tree (What was your maternal grandmother’s name? What was your great-grandmother’s name and date of birth?, etc.).
Here is a very rough schematic of the Reverse Family Tree process:
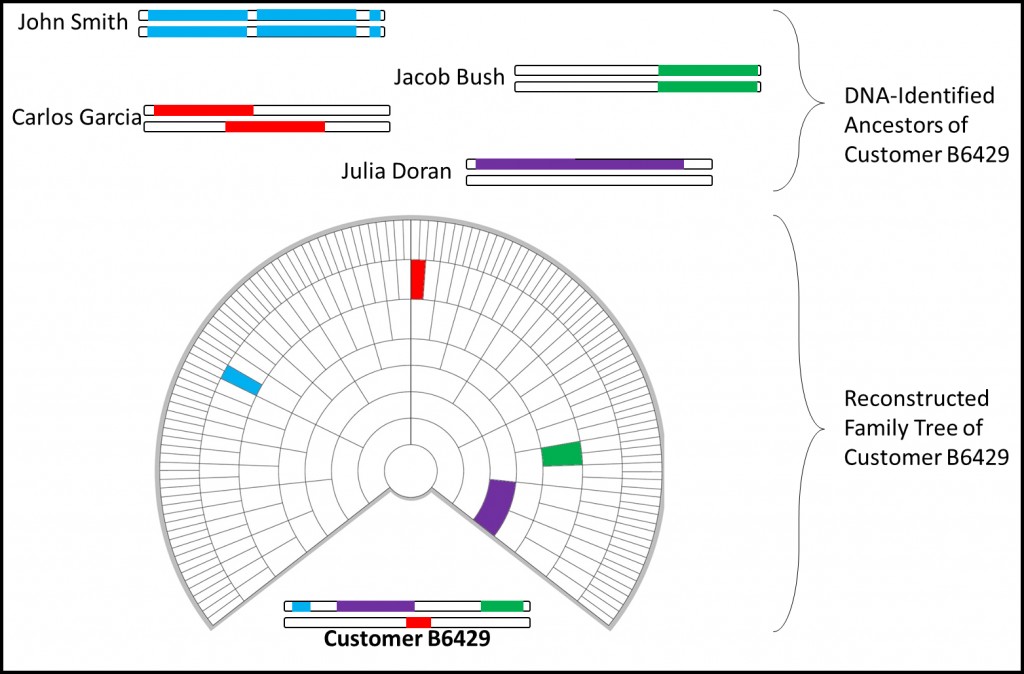
Customer B6429 possesses segments of DNA from four different reconstructed genomes. This information is used to create a Reverse Family Tree with the identified ancestors mapped to it in the most likely configuration based on the size of the segments, established genealogies, and several other factors.
This Reverse Family Tree process (i.e., working forward in time instead of back in time) could also be used with traditional genealogical research, although it would be unlikely to happen. For example, if I were to tell a genealogist only that a client is known to descend from these 35 people, that genealogist could, with enough time and research, recreate the probable family tree of that individual. Rather than recreating a family tree entirely from scratch, the method described above can piece together portions of existing family trees in the database to generate possibilities for the customer.
While the most recent 3 to 5 generations might have to be filled in by the customer (because these generations are the least likely to be included in the company’s database), much of the tree could be completed based only on the DNA results.
While there are many caveats with this method, and any computer-generated family tree should be confirmed by traditional research, it will provide valuable data to customers, especially adoptees.
Lastly, note that these outcomes have been a goal of genealogists for well over a decade, especially the Sorenson family (see “Unlocking DNA: Founder’s goal is to build brotherhood by extending kinship among humanity“) of the Sorenson Molecular Genealogy Foundation.
What About Poor Quality Trees?
As all genealogists know, the world is full of poor quality family trees, both online and offline. While these poor quality trees will impact the method described above, they will not be able to derail it entirely. As more people are tested and more information is collated into the database, the DNA information will overpower these poor trees.
For example, it will become clear with testing that John and Jane Smith of Boston, MA could not have been the parents of Joseph Smith, since none of Joseph’s many descendants possess the unique John and Jane Smith autosomal haplotypes, and instead possess the Samuel and Rachel Smith haplotypes.
In addition to poor quality trees, there are extremely well-researched and well-documented family trees that are incorrect due to an otherwise undetectable NPE (non-paternal event) such as adoption, name change, or infidelity. These trees can also be detected and (delicately!) corrected using the methods described above.
This might also be another motivation for a universal family tree complete with documentation such as the one currently being worked on by thousands of users of FamilySearch.
Other Information
As mentioned above, every ancestor in your family tree with a recreated genome could have his or her own profile filled with information about their life, family, and DNA.
Having the recreated genome of this individual may also provide his descendants insight into other information about him. For example, within that genome may be health propensities (cancer? heart disease?), traits (brown hair? blue eyes?), and other DNA-related information.
It may also be possible to create images of the ancestor based on recent and ongoing research linking facial structure to SNPs within the genome. The DNA information can then be supplemented with cultural and socioeconomic information to predict hair styles and other features.
Caveats
There are many caveats to this process, which is why it cannot completely eclipse traditional genealogical research. Since the blog’s launch in 2007, I’ve stated many times that genetic genealogy only works when paired with traditional genealogy, and that remains true.
For example, it may be that you descend from the relatively unknown brother of John Smith who has very few living descendants, instead of being a descendant of John Smith himself. You could still possess any of the characterized John Smith haplotypes. While the methods described above will likely ultimately focus on characterizing the genomes of “branch points” (such as immigrants, founders of unique haplotypes, etc.) to avoid this problem, the existence of unknown or lesser known relatives of the branch point will temporarily throw a monkey wrench in the process. Many of these issues will be resolved by traditional research.
What are some of the other caveats or issues you see with my envisioned future of genetic genealogy?
I also agree that this is where the future is going. However, I do believe that this technology will need to be evidence based – thus access to the segment data and supporting paper documentation will be imperative.
There are plenty of times that graves are moved and DNA testing is done, over time I’d imagine evidence would surface that way, and should be include into the databases. I would sign off on my ancestors being tested if their graves were ever to be moved. Just one reconstructed DNA profile being matched to a real DNA sample would provide a lot of insight.
What are the public companies (23andme, etc) doing in this regard?
Bill,
The ASHG abstract that triggered this post is from AncestryDNA, so they are currently working on at least a version of this future. 23andMe and Family Tree DNA both allow and encourage linking of family trees to accounts, so perhaps they could pursue this avenue as well.
This would be awesome, especially for African Americans.One major snafu would be privacy issues, how would they get around the private and locked trees, etc.
Earl – I agree, it will be great for many people, as long as there are paper trails to accompany the DNA. As you probably know, this will present an issue for African Americans and other others without well-established genealogies. Also, there is no doubt that privacy will be a huge issue. Thanks for reading and commenting!
This is where chromosome mapping ultimately leads. I hope that many more people begin to explore the value of chromosome mapping, whatever the big DNA testing companies decide to make available. And I hope GEDmatch.com stays open for a very long time to come.
Mini Plus B31: This will be the cheapest deal Keurig offers, featuring three serving sizes and a one-time use reservoir.
For those of us, who for whatever reason, might still
get impatient waiting 10 minutes, it has a Pause-and-Serve feature that permits you to grab your mug of coffee before it can
be done brewing. I must point out that I think personally the brewing temperature ought to
be just over the 200 degree mark for your “perfect” flavor to be extracted, but the Cuisinart does
a solid job.
Thanks for one’s marvelous posting! I truly enjoyed reading it, you will be a great author.I will
be sure to bookmark your blog and will come back sometime soon.
I want to encourage you continue your great posts, have a nice
evening!
When the body is unable to figure out what to do with the
food it may look at it like a toxin and push it off into a fat cell or
rush it through your digestive system as fast as it can to get
it out of there since it is unable to absorb the nutrients which it no longer understands.
Brownies are always yummy, but this brownie dessert tops them all
with the added peanut butter. Allow the pan to
sit for about 10 minutes before serving the oven baked s’mores.
Excellent point to ponder.I am futaonrte to have a set of grandparents I can still call. My grandpa is not a talker and my grandma knows next to nothing about her family. She never knew her mom and didn’t want to bother her dad with questions. Needless to say, it’s been challenging to get information from them. Also, I find that the siblings don’t always keep in touch, are mad at each other, or won’t talk about the past.Thanks for bringing up the issue. It’s been fun to think about the philosophical points..-= Amy Coffinb4s last blog .. =-.