 TribeCode (www.TribeCode.com) is a relatively new direct-to-consumer genetic genealogy testing company, officially launching in the fall of 2014. The company is owned by Centrillion Biosciences, headquartered in Palo Alto, California. The TribeCode test, currently offered for $99, offers Y-DNA, mtDNA, and atDNA analysis.
TribeCode (www.TribeCode.com) is a relatively new direct-to-consumer genetic genealogy testing company, officially launching in the fall of 2014. The company is owned by Centrillion Biosciences, headquartered in Palo Alto, California. The TribeCode test, currently offered for $99, offers Y-DNA, mtDNA, and atDNA analysis.
The ISOGG wiki page about TribeCode offers some information about the test, gleaned mostly from Facebook postings by the company. For example, the test apparently uses an Illumina low-coverage sequencing technology and tests at least 12 million markers throughout the genome. More exact details of the sequencing aren’t yet found on the TribeCode website.
Around Thanksgiving of 2014 I ordered the test on sale from approximately $79, and received my results a couple of months later.
My Ancestry
After receiving notification that your results are ready, you’ll log in and see the introductory page:
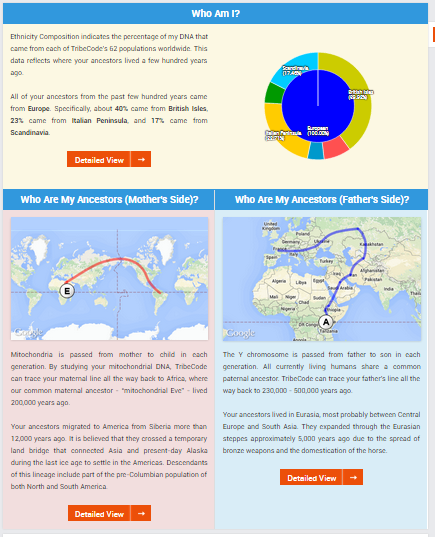
The introductory page provides links to all the various tools, including:
- Ethnicity Composition – Overview
- Maternal lineage (mtDNA)
- Paternal lineage (Y-DNA)
- Ethnicity Composition – European View
- Chromosome Painting (Ethnicity Composition – Chromosome View)
- mtDNA Tree
- Y-DNA Tree
- Jewish Ancestry
- Principal Component Analysis – PCA
I’ll look at most of these tools below, analyzing them in view of tools I’m familiar with at other testing companies.
Maternal Lineage
According to TribeCode’s website, the test “sequences your full mitochondrial DNA, giving you the most comprehensive and accurate maternal lineage information possible.” My mtDNA results show as A2w1, which is the furthest branch of the mtDNA tree that I can currently be mapped to (other than still-new research showing that I am A2w1b, see “An mtDNA Journey – Discovering My mtDNA in a Research Paper“):
For comparison, Family Tree DNA assigns me to the Haplogroup A2w, and 23andMe assigns me to A2. Family Tree DNA’s full mtDNA sequence could classify me as A2w1 and A2w1b, but their tree will have to be updated with the most recent information. 23andMe does not sequence the full mtDNA genome, and a quick look at the SNPs they test indicates that they are probably unable to classify me as A2w1.
TribeCode provides an mtDNA tree interface, which is based on PhyloTree (the most recent version of which is from February 2014):

My branch is very accurate, minus the very recent discovery of new branches under A2w, including my branch of A2w1b.
In the tree, it appears that green means positive for the SNP and red means negative for the SNP. For example, I know that my mtDNA line likely back-mutated C16111T more than 1,500 years ago, and I am negative for that SNP.
Unfortunately, TribeCode does not provide the mtDNA sequence, other than what can be gleaned from the tree.
Paternal Lineage
According to TribeCode’s website, an algorithm called YFitter is used to determine the test-taker’s paternal lineage. YFitter is a statistical tool that can be used, for example, to determine Y-DNA haplogroup using low-coverage Y-DNA sequencing data.
My Y-DNA results, which TribeCode has based on the ISOGG Y-DNA tree, show as R1b1a2a1ac1a2a:
Looking at the ISOGG Y-DNA tree, I see that R1b1a2a1a1c1a2b is R-L1, which I know is my terminal SNP:

TribeCode provides a Y-DNA tree interface with the same green and red indication:

For comparison, Family Tree DNA assigns me to Haplogroup R-L1 (due to SNP testing imported from The Genographic Project), and 23andMe assigns me to R1b1b2a1a1* (an old way to say somewhere below S21/U106).
Unfortunately, TribeCode does not provide the Y-DNA sequence, other than what can be gleaned from the tree.
Ethnicity Composition
The Ethnicity Composition tool compares your DNA to 62 reference populations (I couldn’t find the list), and is intended to reflect where your ancestors lived approximately 500 years ago.
There are two confidence levels, “Confident” and “Standard.” Confident mode shows all ethnicities equaling 3% or more of your overall composition, while Standard mode shows ethnicities equaling 1% or more of your total composition.
At the “Confident” confidence level, my ethnicity is 100% European:
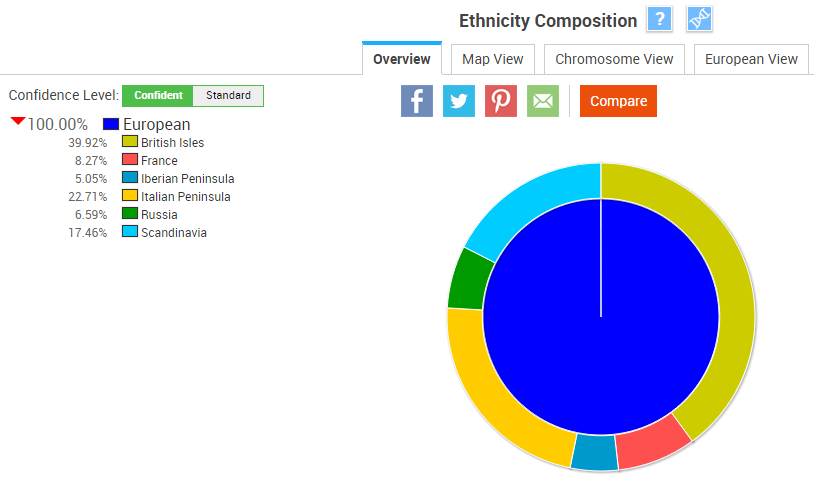
At the “Standard” confidence level, my report adds African, Native American, and East Asian ancestry:
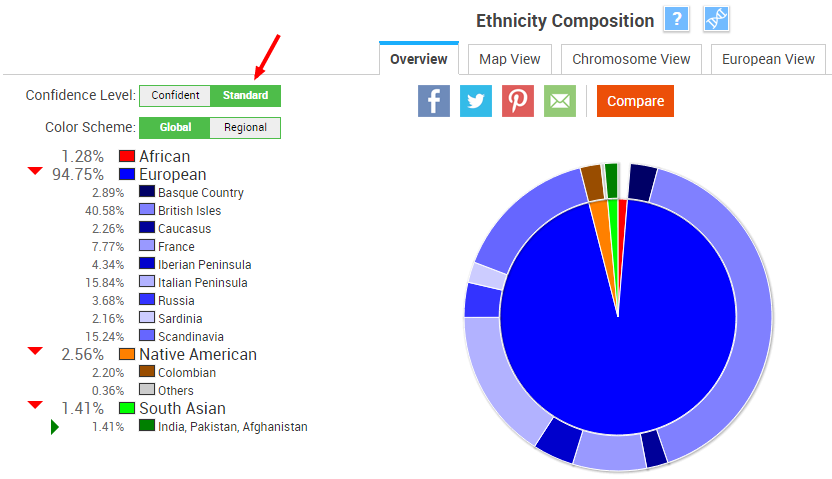
This is very similar to my 23andMe ethnicity report, which is 95.5% European, 3.1% Native American and Asian, and about 1% African (at the “Speculative” confidence level). As we’ll see below, these percentages are not surprising in light of the Chromosome View.
From time to time, TribeCode will introduce “Experimental” tools. For example, a few months ago, the TribeCode ethnicity algorithm was not picking up Native American ancestry, and they had an Experimental tool to detect Native American ancestry. Since that time, it appears that the tool has been incorporated into their standard ethnicity algorithm.
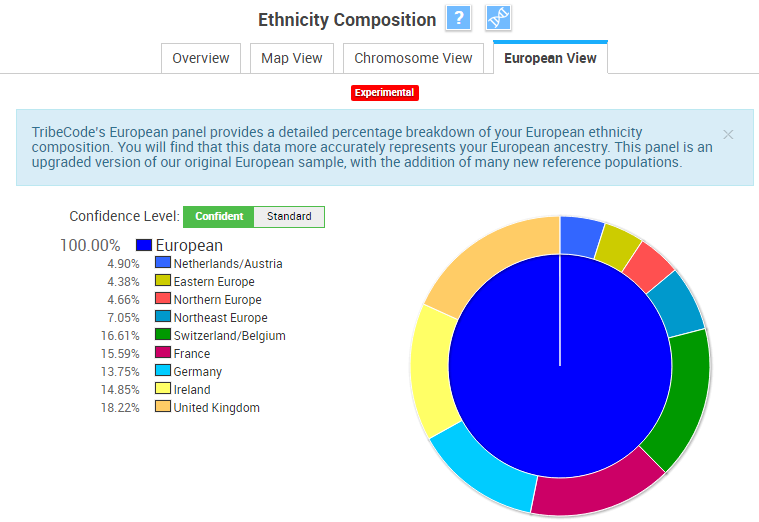
The Experimental “European View” tool shows only European ancestry estimates, and groups all non-European estimates into an “Others” category. In the Standard mode, I have 4.63% Others. It’s interesting to note how different these European estimates are from the European estimates provided above, which is likely a combination of both different labels, and different reference populations.
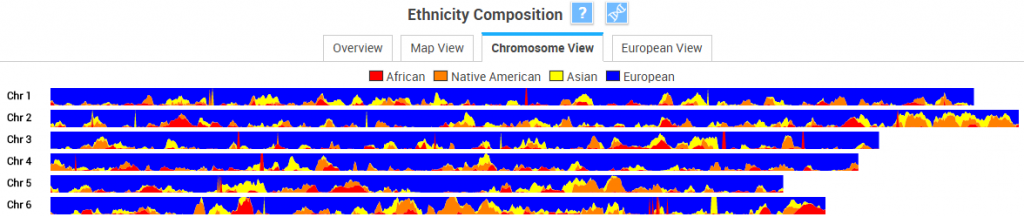
TribeCode also offers a Chromosome View which, like 23andMe and GEDmatch, shows chromosomes 1-22 with your personal calls for the four broadest ethnicity categories painted onto them:
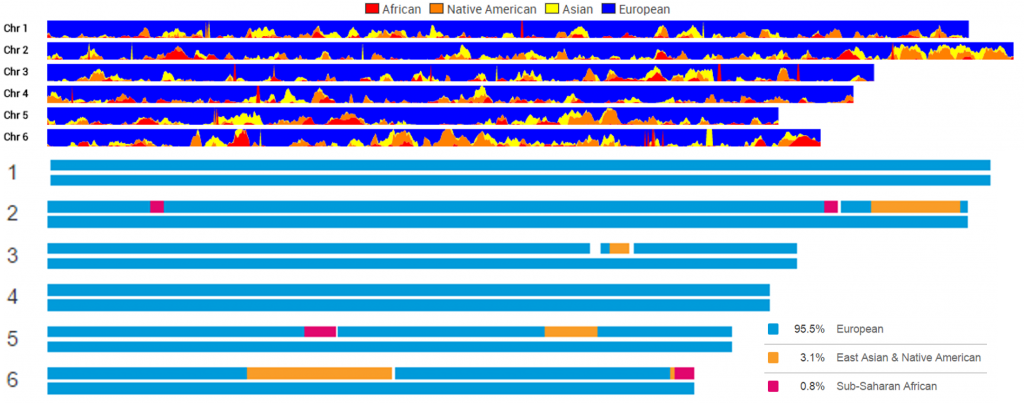
This is a comparison of my first six chromosomes at TribeCode and at 23andMe:
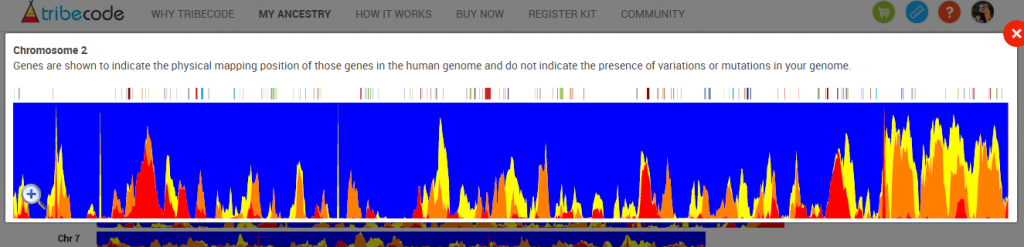
It is possible to look at one chromosome at a time. Here I’m zooming in on Chromosome 2:
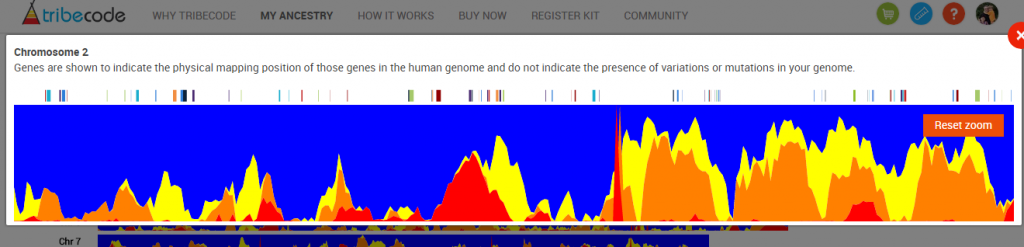
And then you can zoom into a portion of a chromosome. For example, here I’m zooming in on the right end of Chromosome 2:
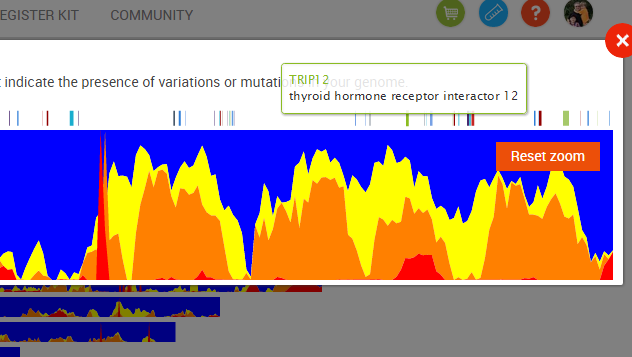
When looking at a single chromosome, you can see the locations of genes along the length of the chromosome (shown by the multi-colored tic marks along the top of the view). Hovering over a tic mark will bring up the name of the gene:
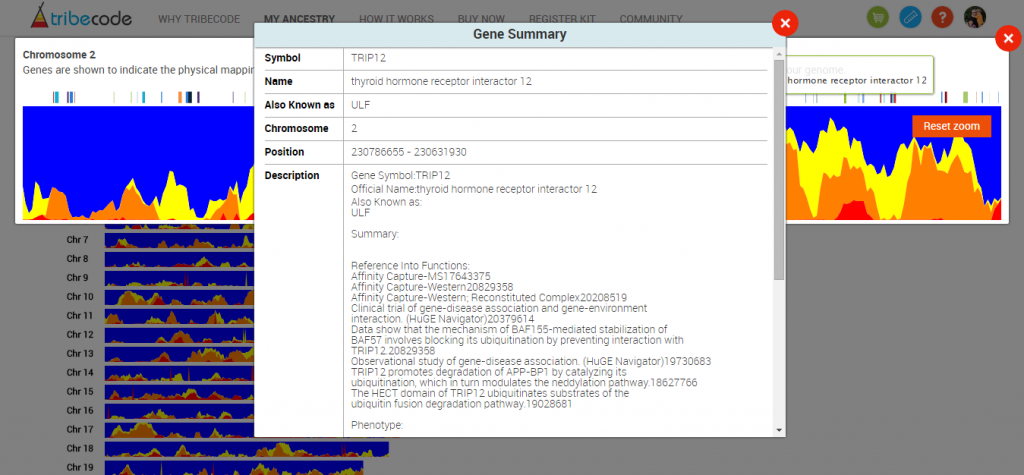
And clicking on the tic mark will bring up a box with information about the gene:
I’m a very big fan of the Chromosome View at 23andMe and GEDmatch, and I very much appreciate having that view at TribeCode. With a few rare exceptions (mostly for adoptees), the overall ethnicity percentages are not very helpful for genealogical research. In contrast, knowing actual segments of DNA that have been assigned to a particular ethnicity could be very helpful to your research.
For example, because I know what grandparent my African and Native American segments come from, I can quickly assign those segments to her. And because I’ve tested my mother, who has additional segments that I don’t have, by process of elimination I can tentatively map those segments I didn’t inherit from Mom to her father, my grandfather. So the Chromosome View at 23andMe helped me map a significant percentage of my genome.
But how do I get the start and stop positions for these segments?
If you’ve tested at 23andMe, you can download a spreadsheet of those segments using DNAGedcom:
You might also be able to get this information from TribeCode’s browser, at least a rough approximation. For example, by clicking on the tic marks denoted by the red arrows below, I can get a gene pop-up with a location.
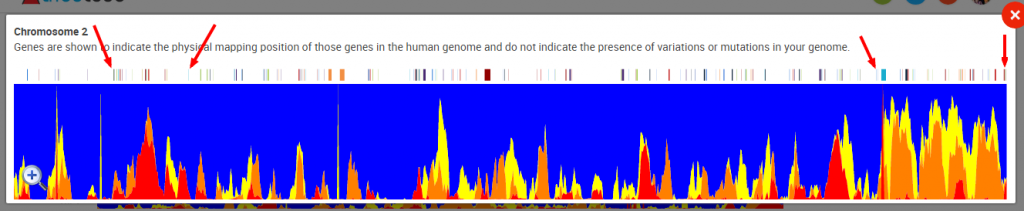
The tic marks reveal the following approximate start and stop locations for these two segments:
- African – 24,397,972 to 42,669,157
- Native American – 211,341,429 to 241,570,676
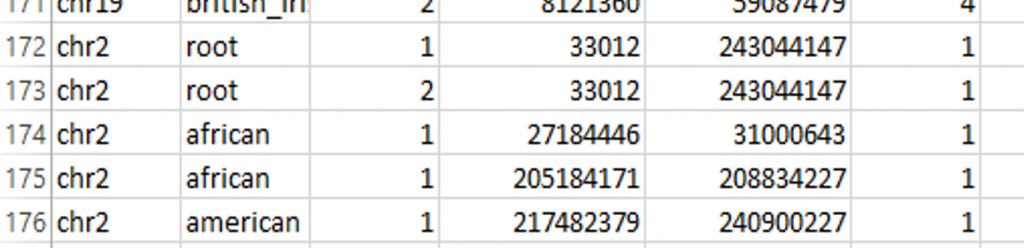
From 23andMe via DNAGedcom:
- African – 27,184,446 to 31,000,643
- Native American – 217,482,379 to 240,900,227
Not very exact, but close. The first challenge, and it is a very big one, is what segments to examine and where to place the start and stop positions. For example, without the benefit of the 23andMe comparison (or perhaps a calculator from GEDmatch using AncestryDNA or FTDNA data), it would be difficult to conclude what segments I thought were significant enough to assign.
Jewish Ancestry
TribeCode also has an experimental tool called “Jewish Ancestry.” According to the website, it can “accurately estimate if you have one, two, three, or four Ashkenazi Jewish grandparents.” From the website:
If you are of European descent, an analysis of your DNA sequencing results may indicate whether you have Ashkenazi Jewish ancestry. Your results are compared to SNP alleles in hundreds of self-described Jewish and non-Jewish individuals from this study, as well as another data set from a large cohort in New York.
I have no known Jewish ancestry:
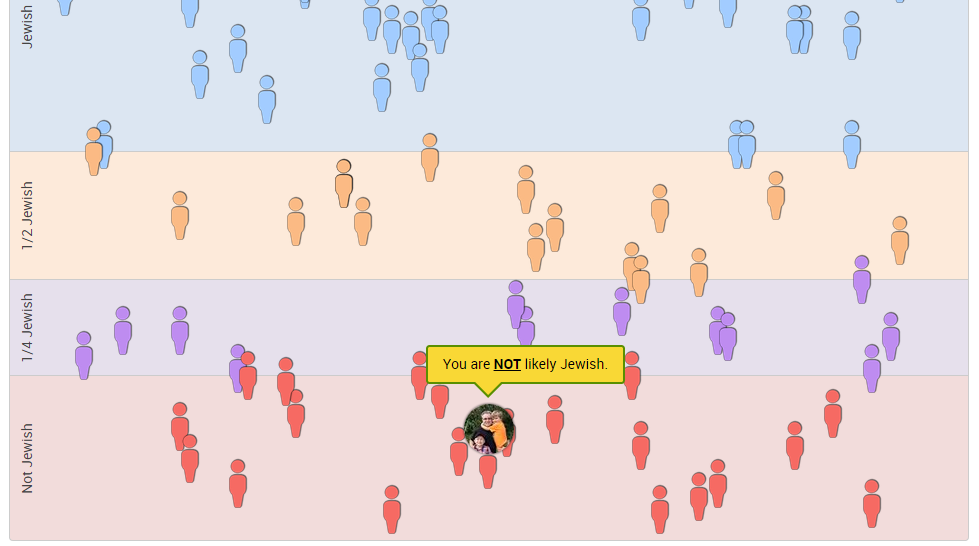
Conclusions
I enjoyed testing with TribeCode and reviewing my results. I think the Y-DNA and mtDNA designations, at least for my results, were extremely accurate. For my Y-DNA, for example, it even determined my terminal SNP.
The ethnicity estimate from TribeCode also appears to be accurate, at least when looking at the four broadest categories. I wouldn’t rely in any way on the sub-regional categories, including the “European View,” but that’s a symptom of the science, not TribeCode. You shouldn’t be relying on sub-regional categories at any testing company.
My concern with TribeCode is that they don’t yet return raw data to customers. According to their Facebook page they’re still considering it, but are concerned about the size of the files which are much larger than typical genetic genealogy files. I’m not positive, but my suspicion is that TribeCode might be concerned about returning low-coverage sequencing to customers (if indeed it is low-coverage sequencing), as it will knowingly contain many sequencing errors due to the low-coverage nature. That doesn’t have a significant impact on the various tools, but it would be difficult to adequately explain to consumers who would find and then only focus on the errors. However, according to Genetic Genealogy Standard #3, genealogists believe that they have an inalienable right to their raw data. Hopefully TribeCode will review their current policy and provide raw data to consumers.
.
I have a few things to say. First, the people who posts insinuating that family members had affairs is disrespectful and irresponsible. Those opinions should be left to yourself. If you’ve experienced that in your family it should stay in your family.
Second, I am concerned about purchasing the ancestry DNA kit but a little more interested in the tribal kit haven’t seen much negative reports.
Third, although expensive maybe getting multiple reports from the three companies 23, ancestry and tribal you might get a little more common denominators.
I’m going to go with tribal and take my chances first.
Keep searching
I tested using Tribe Code and Ancestry. They differ somewhat. Tribe code lists me as 21% Italian Peninsula and 7.5% Basque. Ancestry lists 1% Italian Peninsula and no Iberian Peninsula or Basque. My family tree goes pretty far back and any Iberian or Italian is at least 500 years ago. i seriously doubt their results in these areas.
Back in March, I finally had success with this company after two sample failures. (without an email notification from them, I guess it was because I had my Tribecode account open when they uploaded the results. I guess that kind of bothered me because of my two failures with them even though it shouldn’t because my third sample worked).
Curious why there still isn’t much information about Tribecode online even though Tribecode has had their test since late 2014, which would be over a year and a half now. They told me several months ago that they still haven’t even “officially launched” yet. I think that’s kind of odd that a company that has had a test for more than a year and a half hasn’t “officially launched” yet.
Have people who have received their results obtained reasonable results per individual continents in the TribeCode autosomal test? Just curious; mine were quite believable, but I wanted to hear anecdotal experiences from others. I like their site and how they shared my results with me, but am troubled by the lack of online discussion on the TribeCode test from others, and question whether the company is still viable.
Hi. I think the results in their “confident view” were fine enough. I’m also wondering if this company is even still around and hasn’t folded, because it has now been more than two years since their test has been released and there still hasn’t been much discussion of it online.