NOTE: This is for the (now outdated) June 2016 Update!
[EDIT: PDF edited on 31 July 2016 to correct the averages for 1C and 1C1R (hat tip to Andrew Millard, thank you!)]
The Shared cM Project is a collaborative data collection and analysis project created to understand the ranges of shared centimorgans associated with various known relationships. As of June 2016, total shared cM data for more than 10,000 known relationships has been provided.
This is the first update to the original data, released in May 2015. In this update there are more than 4,000 new entries. Additionally, the data for each relationship has been analyzed statistically to remove extreme outliers and produce a histogram to show the distribution.
For more information see The Shared cM Project page, and “Autosomal DNA Statistics” page at the ISOGG Wiki. HERE is the link to provide your data for future updates.
As always, an enormous THANK YOU to everyone that took the time to provide the data for this project. This is YOUR data!
Known Issues
There are several known issues with the Shared cM Project data, although this update has significantly minimized and/or eliminated these issues:
- Data entry errors – some of the information entered by participants is clearly affected by data entry errors (for example, a longest segment begin greater than the total shared cM). When these errors could be definitively determined, they were removed.
- Incorrect relationships (known or unknown) – some relationships were almost certainly entered incorrectly, which might be due to misunderstandings of “removed” relationships in genealogy. Other relationship errors were clearly due to misattributed parentage events resulting in the believed relationship being incorrect.
The greatest danger posed by these errors is that outliers would significantly affect the maximum and minimum of a range. However, with outliers removed and the histograms provided, these errors are unlikely to have a significant effect on the ranges in the project.
The Data
This update utilizes a total of 9,417 submissions. Although more than 10,000 people have submitted data, this update began a few months ago, and some data points were removed as outliers.
For the first time, the Shared cM Project data also it also contains data from relationships at AncestryDNA. The following graph, for example, shows the bump in submissions in November 2015 when AncestryDNA began sharing that data and I sent out a request for submissions:
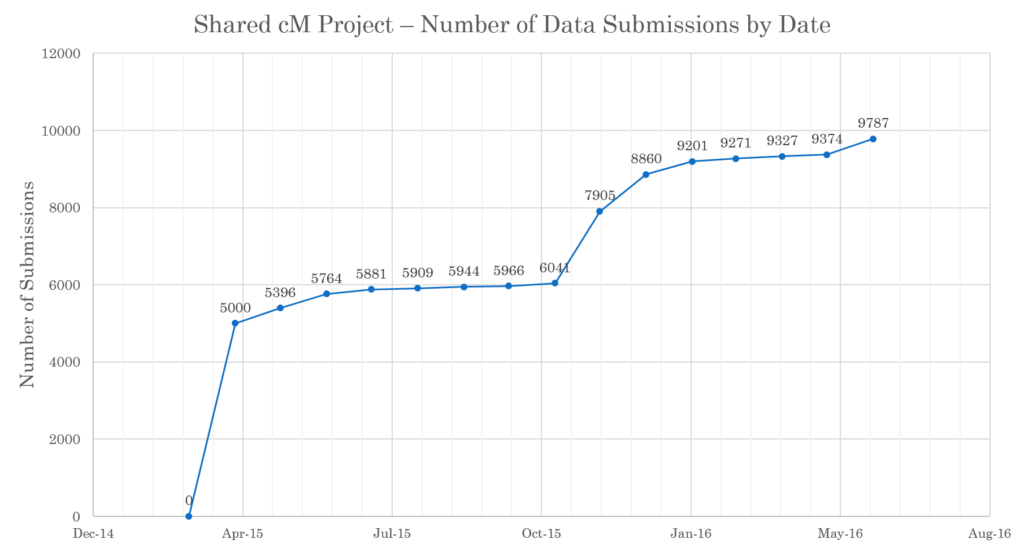
An updated graphic is provided, which includes many more relationships, out to 8C (although be sure to note the low number of entries for many of the distant relationships).
Histograms
This update provides histograms, or distributions of the data entries for each relationship. The histogram for each relationship shows more clearly how your relationship compares to everyone else that submitted data for that relationship.
The file with histograms and other information (including ranges for additional relationships not shown in the chart above) is here: Shared-cM-Project-Version-2-UPDATED (PDF) (Updated 1C and 1C1R averages on 7/31/2016).
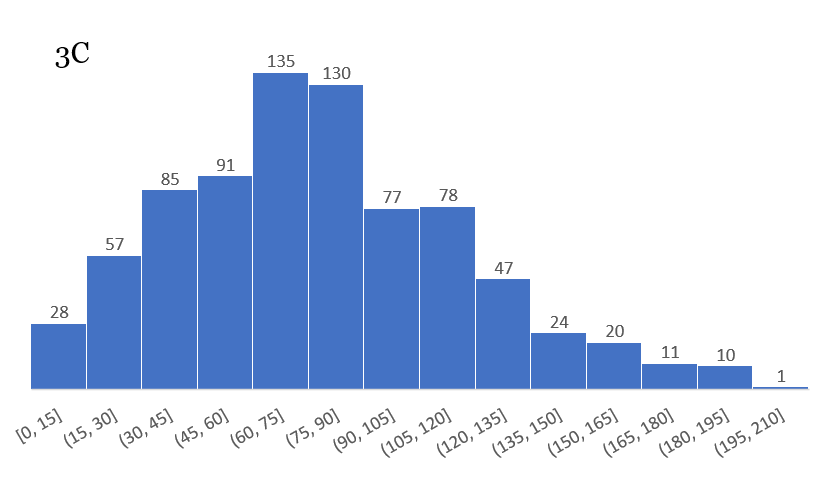
Some of the histograms are truly incredible, showing a perfect distribution:
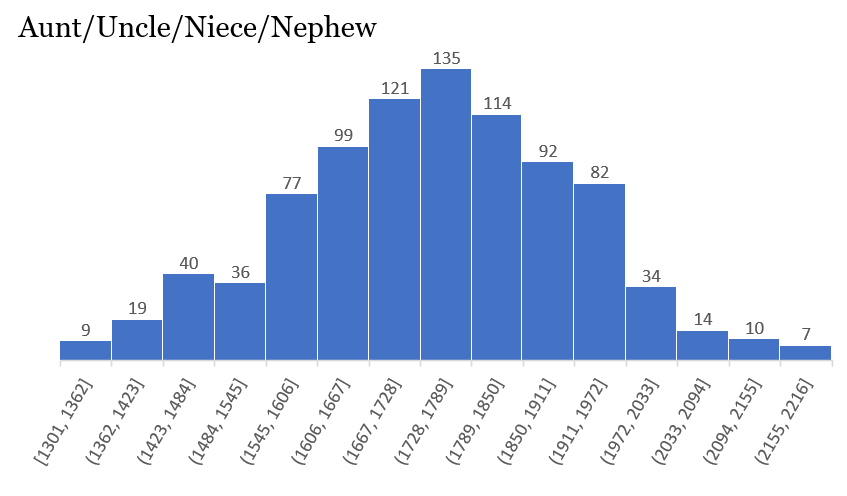
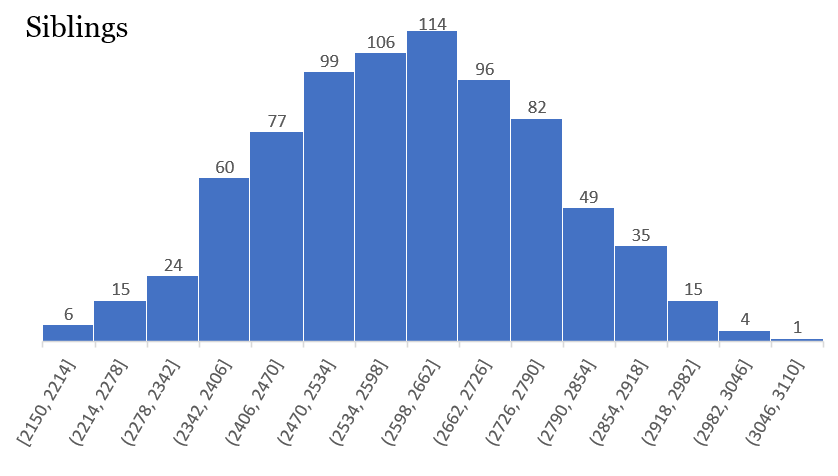
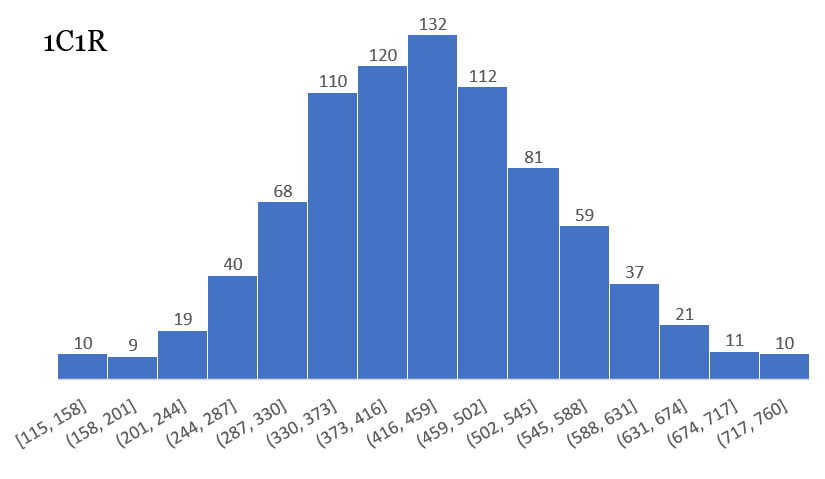 Others are not quite as perfect, but contain a wealth of information:
Others are not quite as perfect, but contain a wealth of information:
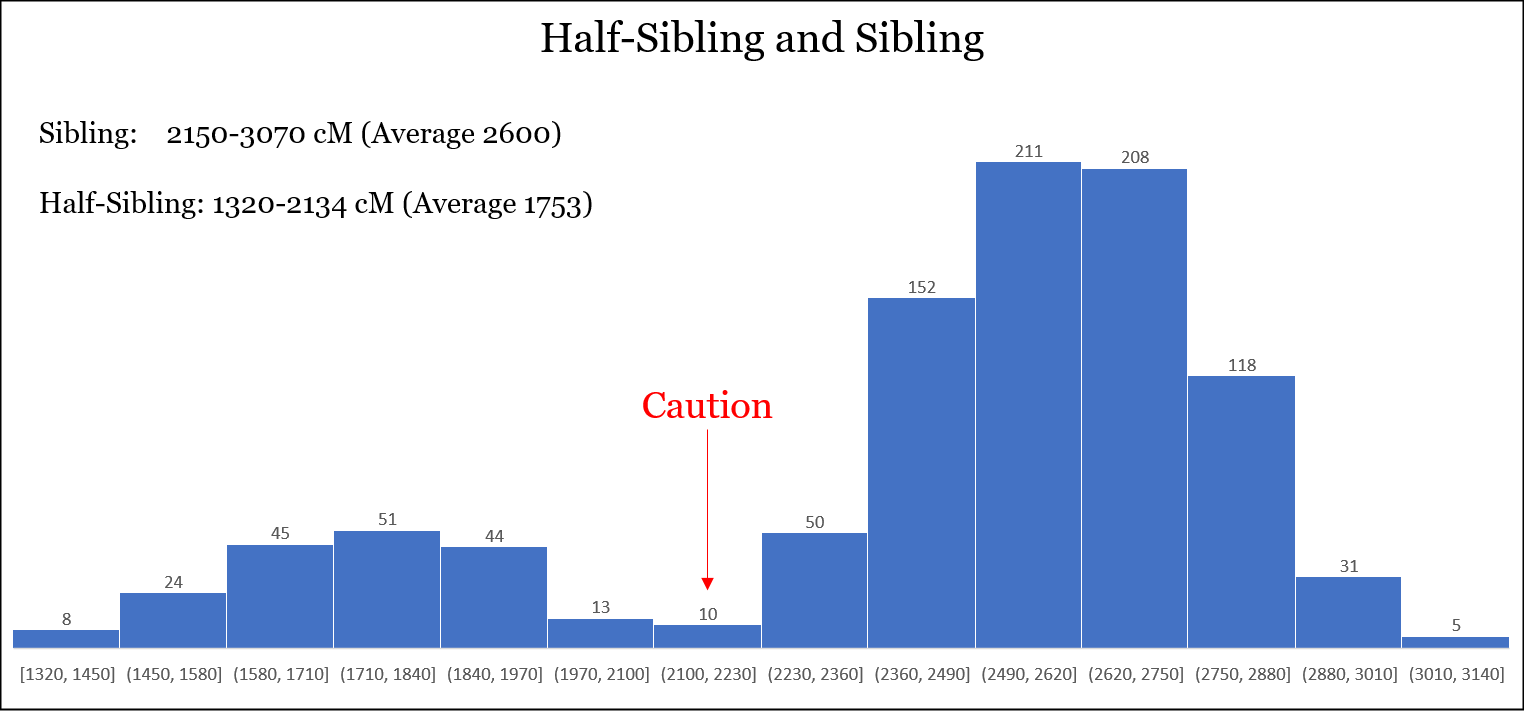
 The following is a histogram containing the data for both siblings and half-siblings. Although the outlier-removed ranges do not overlap, the difference between the highest sharing for half-siblings and the lowest sharing for siblings is about 15 cM. So there is the potential for overlap in that bin, according to the data:
The following is a histogram containing the data for both siblings and half-siblings. Although the outlier-removed ranges do not overlap, the difference between the highest sharing for half-siblings and the lowest sharing for siblings is about 15 cM. So there is the potential for overlap in that bin, according to the data:
For ALL the histograms and ranges, be sure to download the full file: Shared-cM-Project-Version-2-UPDATED (PDF).
Love your work Blaine.
I will be adding the ‘notional’ or ‘expected’ cM to the chart as a point of comparison.
This is an extremely helpful piece of work.
Cheers
Mike
Thank you Mike!
My paternal uncle and I share 1931.01 which falls right in your parameters. However, my uncle and his great-niece (not my daughter but rather the daughter of one of my first cousins) comes in at 1144.32 which is quite high.
Comparing me to her (she would be my 1st cousin, once removed) we share 504.47 which again, is right on target.
It is interesting that my uncle and this great-niece share so much DNA. I wish I could test her father (my first cousin) but he is under the radar and doesn’t want to be found. His father (my uncle’s brother) would rather eat roach droppings than take a DNA test.
I think that if I had my uncle’s brother’s DNA and his son’s DNA it would make an interesting study so determine the big hunk of shared DNA between my uncle and this particular great-niece.
It would be interesting to test your first cousin! Maybe someday!
A great update Blaine – this will be the “go-to” chart for many of us. Thanks for staying with this project.
Thank you so much Jim!
Darn, now I have to print and laminate a new Chart! But Thanks anyway!
Are you still taking submissions? I have several to add.
Haha, sorry about the new chart! 🙂
I’m still accepting submissions. Here is the link to provide your data for future updates: https://thegeneticgenealogist.com/2015/03/04/collecting-sharing-information-for-known-relationships/
A suggestion: It might help to have a footnote or more explanation about “Outliers” and what that means for those of us who have that type of data that was not included. That might provide some help for other “outliers” that might show up in the future.
Kudos: Thanks for all your work on this project. It has been a huge help 🙂
Thank you for the kind words Barbara!
It’s tough to include every possible scenario to explain “outliers,” but it is a great idea to provide some explanations. I don’t have much time to do it right now, but if you and/or anyone else wants to write up a paragraph or two to explain what outliers might be, I’d be happy to review it and possibly add it to the blog post!
One of the striking things is that the average cM is, in all cases, greater than the calculated/expected cM. This is particularly the case in the more distant cousins.
Perhaps, if we are to inherit any DNA at all, it will be more than expected? Or maybe the database contains plenty of endogamous relationships?
This is causing me to rethink some of my more distant cousin relationships where I know there is some endogamy, and others where there isn’t.
If nothing else, questions like these demonstrate how little we really know, and how much more study is needed!
We need SO MANY other citizen science projects like this one to analyze data.
Blaine and Mike,
There’s a logical reason why the average cM are greater than the expected cM, and this is because of sampling bias. For the more distant relationships the only way people discover that they are actually related is because of shared DNA. There are certainly many distant relationships that remain undiscovered because there is no shared DNA or the amount does not meet the companies matching threshold. By its very nature your database can only contain information for relationships that have been determined and for distant cousins these will generally be ones with above average shared cM and will exclude those with lower than average shared amounts.
Here’s an example. So far I have only found one family tree connection among my father’s matches on FTDNA. That match is to a sixth cousin and they share 40.16 cM, well above the expected average of 0.83 cM. But there could be 99 other sixth cousins who have tested but aren’t showing up on his list of matches. Let’s say that those other 99 cousins match my father with a total of 43 shared cM between them. That’s a grand total of 83 cM with 100 sixth cousins (average of 0.83 cM) but the only data that would get entered into the Shared cM Project is the one match of 40.16 cM.
Note that I’m not suggesting any deliberate bias in the reporting or processing of the data, it’s just that the method of gathering data will naturally be biased towards the higher than average shared amounts.
Good point Philip.
Blaine
Really appreciate your comprehensive work in collecting, analyzing, and developing the aggregated data. Very useful.
Two questions:
1) are you going to provided an analysis of the differences between the various versions of test results (23andMe v2/v3 vs. 23andMe v4 vs ancestry v2 vs ancestry v3 vs FTDNA)? As we saw with the analysis done by DNAadoption for our prediction chart, there were noticible differences between the ancestry data and the non-ancestry data.
2) is anyone doing analysis to see the impact on matching (or lack of matching) based on the differences between the chips and algorithms used by the various companies?
Richard,
Thank you for the kind words! Regarding your questions:
I didn’t collect information about test versions, only about testing companies. I would like to break all the data down based on source (23andMe vs. FTDNA vs. AncestryDNA vs. GEDmatch), which is as close to test versions as I can get with this data. Company thresholds and algorithms like Timber are likely to have an effect on total shared cM, as are differences in how each company calculates cMs.
The company differences (particularly Timber) are unlikely to have a significant effect for close relationships (where “significant effect” would mean a clear shift from one relationship to another using the the ranges and histograms herein), but I predict that they could have a larger effect on more distant relationships.
I’m not aware of anyone doing specific studies looking at impact on matching between chips and algorithms at the various companies, although there are usually blog posts discussing matching changes after any major algorithm implementation or change.
Hi Blaine,
I have a question about determining half siblings. We have been searching for my husband’s father. The person we think is the father has passed away but we have located a son of his. My husband did his dna through ancestry and also uploaded it to gedmatch. The other person just recently sent his dna into ancestry and we are waiting on the results to come back. The dilemma is the person that we think is the father and his father had the same name. So we are not sure if my husband is a half sibling to the son tested or if my husband would actually be his uncle. I know a half siblings share a average of 1753 cM. What would my husband share with the other person if he is a uncle. Would it be 1744cM or would he be considered a half uncle.
Renae – if the person you think is the father, and the other test-taker’s father, were full brothers, then your husband would be a full uncle to the other test-taker. Unfortunately, you see that the ranges for half-sibling and full uncle are almost identical, amazingly so. So the atDNA result is likely to narrow it down to the two possibilities (or neither, which is always a possibility!).
Very helpful, as usual!
Blaine, I couldn’t find any instruction on this, but what are your wishes regarding permission to share the chart? I chair a DNA Special Interest Group for our local genealogical society. May I print and share with our group, with proper attribution to you and link back to your blog? What are your preferences on this?
Jacqi – the data is shared under a Creative Commons 4.0 Attribution license. So please print and share away, all I ask is attribution!
Hi Blaine,
Thanks for leading this effort. One question is whether we can generally treat these as additive when more than one pathway is involved? For example, I have one line that likely involves a 5th cousin relationship + a 6th cousin relationship.
Second question, do you know whether Ancestry report any matching X DNA in their shared cM totals?
Third question, I thought that AncestryDNA includes some Y SNPs at some point. Do you know whether those contribute significant amounts of cM to the results.
Finally, it seems that we should expect the “actual” amounts shared to be higher than expected, because of reporting bias. I wonder whether it would be useful to separately report results for tests when one didn’t know the results in advance.
Thanks!
Jonathan
I have a father – daughter relationship I would like to add but the daughter has only tested on 23andme while her father has tested at FtDNA and Ancestry. How would you like to handle this?
Rosemary
I have a cousin with the surname Stills. But we are not related by the Stills Surname. Her William Henry Stills raised my Smith Alexander Ricker as a Stills with his children; from which she descends.
Through Ancestry testing, I am related to her 6 different ways through her mother (a Ricker) and her father. These show as being in the 4-6 cousin range. I am related to her though both my paternal grandmother and grandfather, who were 3rd cousins.
Gedmatch says she shares 159.6 cM’s with my Father and 73.3 cM’s with me.
My goal is to prove Smith Alexander (Ricker) Stills is truly a Ricker. Lots of circumstantial evidence but I was hoping to use DNA to confirm.
(I am still fishing for a Y-DNA match.)
I am learning but I think this is one to leave alone for now.
Thank you for your tremendous resource! I will be sharing your chart (and the link to your ebook) at my upcoming program on DNA for the Bucks County Genealogical Society. I took your course at GRIP last summer and now mentor others in utilizing this invaluable tool for genealogy. Finding an unknown distant cousin through DNA was so beneficial to me because she had traveled to the European homeland and obtained the church records! This furthered my tree two more generations and countless branches, which then helped me determine the MRCA with previous matches.
I really appreciated the shared DNA chart from the SharedcMProject. By any chance is there a rule of thumb or calculation that can be generally applied to the average amount or range of amounts for various relationships that will provide an idea of how much cms of DNA would be shared if those relationships were half-relationships? I am suspecting that it would be way easy to simply divide the amount in half.
My mom shares 537 cMs with her first cousin twice removed and I share 221 with the same person (for me 1st cousin, 3x removed). Is this unusual?
Blaine, I am so thankful that you have created this chart and keep updating the data in it as more and more data becomes available.
PS. speaking of data entry errors, I read and re-read this sentence in the article above wondering why the beginning point of the shared segment was relevant, until I realized you meant “being” instead of “begin”. (for example, a longest segment begin greater than the total shared cM)
Hi Blaine,
Thanks for your work. I use that chart as a default template to calculate relationship range more clear comparing to FTDNA. But I faced with dilemma, that I can’t figure out a few items:
1) Can this chart be applied for FTDNA data?
(I’ve read above thread/comments, but still not sure if this chart u’ve done reflect dedicated database of data or widely used data sources like ftdna, 23andme, ancestry.com, etc.)
2) I do compare atDNA matches for me, my father and mother, and sometimes I have common matches with mother or with father or with both. And you might guess, FTDNA defines relationship range differently. And it’s expected. But sometimes, I have more shared cM than my father with some matched person. And as result the chart I use, give me at least 2 variants of calculation.
Example: me, my father, Match A (no official relation, only DNA shared segments).
– me and Mach A – 36cM (12 shared segments, 1cM threshold, Chromosome browser)
– my father and Match A – 26cM (8 shared segments, 1cM threshold, Chromosome browser)
I take into consideration avg value from your chart, so
– 36cM is 3C2R or 3C1R and 4C.
– 26cM is between 4C and 4C1R.
So question is – how to define the relation in proper way, when I see a few variants, based on your chart?
Note: I’m not professional geneticist at all (I’m web developer, genetics is my hobby), so plz excuse me if I am not correct somehow.
Dear Blaine,
I have recently had my DNA results back from Ancestry.
Although I live in Sheffield , I am a member of the OFHS and found the recent article Vol 30, No 3 by Richard Merry very interesting.
My paternal side comes from Haddenham, Bucks, but have connections with Oxfordshire
Would it be helpful if I submitted my DNA results for your project.
Best Regards
Rodney Slatter
Please forgive me this is way above my head. I just recently did the DNA test and I am finding results that say I have 4th to 5th cousins that have done this as well. In reading this chart the I am thinking these people would be in the 3C to 3C1R area which would mean we have 2nd great grandparents in common. I am I reading this correct? How could I tell looking at the dna data provided to me if it is paternal or maternal? Thanks anyone for responding.
I am pretty new to this and like Kevin in the above message, this is a little over my head. I have a match that has 197 shared CMs with me. Do I find the square that has the closest average to the 197, i.e. in this case I see three possibilities: 1C2R, 2C, and 1C2R, is that right? And, this has me starting to look at this individual’s grandparents, 1x great-grandparents, and 2x great-grandparents to find our common ancestor?
Also, this chart is for cMs, so this connection could be through my paternal and/or maternal lines?
Thanks in advance!
Does the input for the form include X chromosome? Or just 1-22 chromosomes?
I recently have an autosomal dna match with a female(Subject A) and we share 997cMs on 38 segments at ancestry. We have communicated and according to our genealogy we are 1st cousins 1x removed to each other. Subject A’s great-grandmother was my maternal grandmother. Our shared cMs seem more like a 1st cousin; however I have a theory of why we match so much dna. My birth mother was the older sister of Subject A’s grandmother, and both of these sisters shared the same paternal family line. My birth father & birth mother shared the same paternal great-grandfather, so they were 2nd cousins. So I believe my paternal dna outweighs my maternal dna, which might explain why I share so much dna with my 1st cousin 1x removed?
Best regards, Doug
I read your post and wished I’d written it
Thanks for reading our blog. Both Latisse and Lumigan are by picterrpsion. They both have the same active ingredient, however Latisse comes with special brushes that allow you to apply it to the lid margin. I really don’t know if it is available in Singapore. I would recommend checking with your eye doctor. Repeated application of Latisse to any skin tissue is reported to cause hair growth. The most common side effect Latisse is increased pigment in the skin, however Allergan reports that the hyperpigmentation goes away when Latisse is discontinued. I hope this helps. Good luck.
Do you have any idea what half-siblings whose fathers were brothers would look like? I have some odd results between three girls who are supposed to be full siblings. It is obvious that that is not the case. But one rumor involved the mother sleeping with the father’s brother and he fathering one of the children. Looking at gedmatch, I don’t think that is the case. I would assume they would share more cMs than the average half-siblings, which two of the sisters do (2042 cM), but they do not have any significant fully identical regions. They do have more green segments than when comparing them with the third sister, but I still don’t consider them significant. But then again, I am uncertain if they would have fully identical segments and, if so, how large those segments would be, if the fathers were brothers. It is so confusing. It sounds like the start to a bad country song. But I was hoping you might know what that would look like on gedmatch?
Also possibly of significance, I ran the kits through the “Are Your Parents Related Tool” and out of the two sisters who share 2042 cMs, one kit comes back as not related, the other comes back as related within 3.4 generations. I am not certain how accurate the estimation is though, because the mother comes from an endogamous population, and I guess potentially the father does, too.
Oh dear Pip. Being gay is not a disease. It can´t be caught. In affairs of the heart, one should be free to demonstrate their love in whatever way they see fit. These two guys wanting to commit to each other does not in any way affect you or anyone else. Love between two adults, no matter their sexual preference should be celebrated and not damned. By the way Pip, I´m gay.That is who I am and I refuse to hide myself from the world to protect those that feel threatened by it. Get over it!
My mom was told her father on her birth certificate was really her uncle and her uncle(her fathers brother) is her father ( her mom and husbands brother had an affair). Anyways my mom took a DNA and matched ‘close family’ to a lady who should be her cousin (her uncles daughter). But the centimorgans were 1, 406. I realize its not as high as the average but is high enough to be half siblings. How can this be determined ? Isn’t a 1st cousin going to be under 1,000 centimorgans
Of course the closer relationships would show the typical Gaussian curve distribution, and chance in the “DNA Roulette” of more distant relationships as recombination plays a larger and larger role starts shifting the curve. The only relationships being counted are the ones where there actually is a match, and the chances of not matching at all grow with every generation.
Interesting that I match my uncle at 1990 cM, which appears to be out in the “two standard deviation” range of the distribution curve, but obviously we are not alone there. Seeing that makes me tend to think more seriously about using his DNA for phasing since I cannot get my mother’s, but I do have my Dad’s.
On the other hand I match my 1C1R at 377 cM just off the median.
I use these charts regularly when working through DNA matches and have found them to be incredibly reliable and immensely useful, (it helps to understand the statistical methods being used.) And understanding that outliers do exist, and that they have been removed from the equation, helps as well.
Thank you for your hard work.
In searching for my birth parents, I have a closest match sharing 2,220 cm across 66 segments. WE’ve narrowed down a ‘half uncle’ to him that could be my birth father, and that would make this person my 1st cousin. But we share SO much DNA… almost like half siblings. Can 1st cousins ‘half’ share this much DNA (2,220)? My daughter and her half sister only share 1,300cm.
Ok Boomer