For reference, here are all posts for the Shared cM Project:
- Most up-to-date post: “Version 4.0! March 2020 Update to the Shared cM Project!” (March 27, 2020)
Older Posts:
For reference, here are all posts for the Shared cM Project:
Older Posts:
Interested in learning about the unique inheritance of the X chromosome through the use of some cool visual charts? Look no further:
And be sure to check out Debbie Parker Wayne’s great charts at “X-DNA Inheritance Charts“!
Have you experienced this? You’ve identified a very clear cluster that includes numerous DNA matches that all descend from a single family, but you have no idea how this family links into your family tree. Try as you might, and despite building numerous trees, you can’t seem to figure out how these DNA matches and this single ancestral family link into your family tree. If this sounds familiar, you have an Unlinked Family Cluster!
Defining “Unlinked Family Cluster”
An Unlinked Family Cluster is a very specific phenomenon in genetic genealogy, one that is becoming increasingly common. We see more and more of these clusters for various reasons; the matching databases get larger and larger meaning that these clusters get larger and easier to identify. Additionally, the more we work with our closest DNA matches, the more we have very promising “left behind” matches that don’t fit into our known ancestry. Often, these “left behind” matches form a shared match cluster or a triangulated cluster around a specific family. This is an Unlinked Family Cluster.
The Shared cM Project is a collaborative research project that uses data from 100s of genealogists to generate shared DNA ranges and averages for nearly 50 different genealogical relationships. The most recent version of the Shared cM Project, Version 4.0, was released on March 27, 2020 (I had so much free time in the early days of the pandemic!). See “Version 4.0! March 2020 Update to the Shared cM Project!” Keep those submissions coming, I’m hoping to do an update in 2023!
Always Improving!
One of the enhancements to the Shared cM Project that has been on our minds since the creation of the project is the use of siblings and/or same-generation cousins to narrow down the list of possible relationships to a DNA match.
For example, my sister and are full siblings, and thus by definition we have the same genealogical relationship to all of our genealogical cousins. Sometimes this can be a challenging concept! This graphic shows how my sister (Sibling #1) and I (Sibling #2) have exactly the same genealogical relationship to this third cousin (3C):
The Shared cM Project is a collaborative research project that uses data from 100s of genealogists to generate shared DNA ranges and averages for nearly 50 different genealogical relationships. The most recent version of the Shared cM Project, Version 4.0, was released on March 27, 2020 (I had so much free time in the early days of the pandemic!). See “Version 4.0! March 2020 Update to the Shared cM Project!”
Several years ago, Jonny Perl – creator of the invaluable genealogical website DNAPainter.com – graciously converted the results of the Shared cM Project into a dynamic online tool. With this FREE tool, a genealogist can enter an amount of DNA shared (in cM) with a match at a testing company, and the possible relationships for that shared amount will highlight in the graphic. In addition, clicking on any of the relationship boxes produces a pop-up that reveals the histogram for that relationship (arguably the most valuable aspect of the Shared cM Project).
TL:DR Executive Summary:
Small segments have long been a controversial subject in the genealogy community. Some love them, some hate them. Here we will look at all the available evidence surrounding small segments, some of the misconceptions associated with them, and some of the ways we might be able to utilize small segments in our research.
One of the biggest problems surrounding the discussion of small segments it that there is no strict definition of the term. A “small segment” to one person might be anything 5 cM or less, while a “small segment” to another person might be anything less than 10 cM. And there are many other variations. For purposes of this analysis, we will define a small segment as any single segment of DNA less than 8 cM (although segments as high as 10 cM or more can be problematic as well).
During a phone call with AncestryDNA representatives this week (unfortunately I was not able to attend), numerous genealogists heard two major announcements:
The announcement started to appear on the DNA match list page yesterday:
Clicking on the link brings up information about the changes:
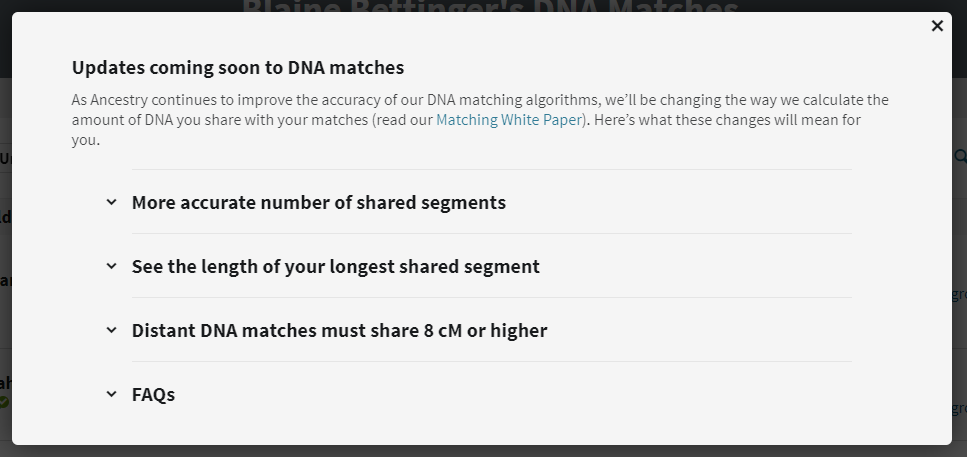
The changes to the DNA match list comprise the following:
From the announcement: “The DNA you share with a match is distributed across segments – short segments, long segments, or some combination of both. Our updated matching algorithm may reduce the estimated number of segments you share with some of your DNA matches. This doesn’t change the estimated total amount of shared DNA (measured in centimorgans/cM) or the predicted relationship to your matches.”
The Shared cM Project (ScP) is a collaborative data collection and analysis project created to understand the ranges of shared cM associated with various known relationships. The ScP has been very successful, with more than 60,000 submissions from amazing genealogists like YOU! To add your data, the Submission Portal is HERE. I am always collecting data, and hopefully the next update will have more than 100,000 submissions!
The full PDF for Version 4.0 of the Shared cM Project is here and it is ESSENTIAL that you read the full PDF for all the details from the project: The Shared cM Project Version 4.0 (March 2020).
Today, the most recent version of the ScP, Version 4.0, goes live. I’ve taken nearly 60,000 submissions and analyzed the data for almost 50 different relationships. For each relationship the 100s or 1000s of submissions were analyzed to remove outliers, to provide minimum, maximum, average, and standard deviation values, and to generate a histogram for the distribution of the submissions. Here are some of the other differences between this new Version 4.0 and the previous version (click to enlarge):

[WARNING: I discuss or imply violent acts by ancestors in this post, read at your own risk].
We’ve all heard it. Some of us have even made it. A joke or implication about an affair or dalliance that conceived a child, often referring to the milkman or a neighbor. It’s usually directed to the biological mother, always ignoring or downplaying any act by the biological father, and is always consensual. The audience (whether in a Facebook forum or at a talk/seminar/webinar), seemingly always primed for the joke, laughs and the speaker moves on.
It’s time for this joke or implication, whether blatant or implied, to die the ignoble death it deserves.
An Admission
A few years ago during a lecture, I make a flippant remark about a misattributed parentage conception. It may have been as simple as raising my eyebrows at a key moment, or even a simple pause that implied meaning, I don’t remember. After the talk, an audience member came up and called me out for being flippant about misattributed parentage conceptions. And the audience member was right, I had been flippant. I was wrong.
In this blog post we will briefly review an extreme Grandparent/Grandchild relationship, where a grandchild appears to share just 9% of her DNA with a paternal grandmother rather than the expected 25%. All information is anonymized.
I’m a little afraid to post this article about an extreme outlier scenario. There is a danger that it could support misinterpretation rather than foster critical thinking. If you have a possible outlier scenario, be sure to try to disprove that it is an outlier situation, rather than simply proceeding as if is an outlier. Avoid confirmation bias!
This is the third post on my blog specifically examining outliers in confirmed relationships:
This was discussed in a previous article about outliers, but it bears repeating.
The annual RootsTech convention at Salt Lake City in February has become a showcase for new tests and tools offered by the DNA testing companies. The biggest winner of all, of course, is the consumer!
Both AncestryDNA and MyHeritage announced major new developments at this year’s RootsTech. For example, AncestryDNA announced “MyTreeTags,” “New & Improved DNA Matches,” and “ThruLines,” three different tools currently in beta. The first two require an opt-in, while everyone is currently eligible for ThruLines. Meanwhile, MyHeritage announced “Theory of Family Relativity” and “AutoCluster,” both of which are currently available to members of their DNA database.
There is a LOT to digest with these tools, including knowing how to use them and understanding their limitations. To help you understand what the tools are and how to use them, I’ve created two new YouTube videos:
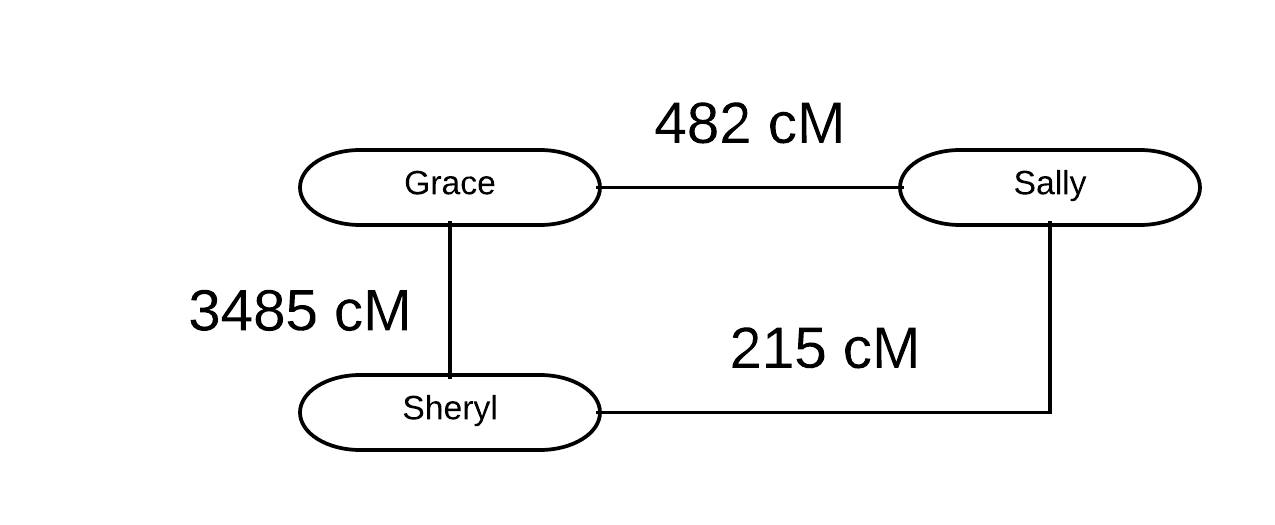
Sheryl, a member of the Genetic Genealogy Tips & Techniques group (which just broke 50,000 members!) recently commented on a thread about shared DNA outliers about a situation within her own family. I thought it would be a great opportunity to discuss outliers and how to deal with them. Sheryl kindly agreed!
For background, we examined an outlier situation once before on this blog, where second cousins once removed (2C1R) did not share DNA (see “Analyzing a Lack of Sharing in 2C1R Relationship“).
Sheryl indicated that she and her mother Grace appeared to be outliers with Sally, their first cousin (1C) and first cousin once removed (1C1R), respectively. Grace shared 482 cM with her 1C Sally, and Sheryl shared 215 cM with her 1C1R Sally. Not surprisingly, Grace and Sheryl share an expected amount for mother/daughter: