During a phone call with AncestryDNA representatives this week (unfortunately I was not able to attend), numerous genealogists heard two major announcements:
- The AncestryDNA database has hit 18 million test takers (such great news!); and
- There are significant changes coming to our DNA match list.
The announcement started to appear on the DNA match list page yesterday:
Clicking on the link brings up information about the changes:
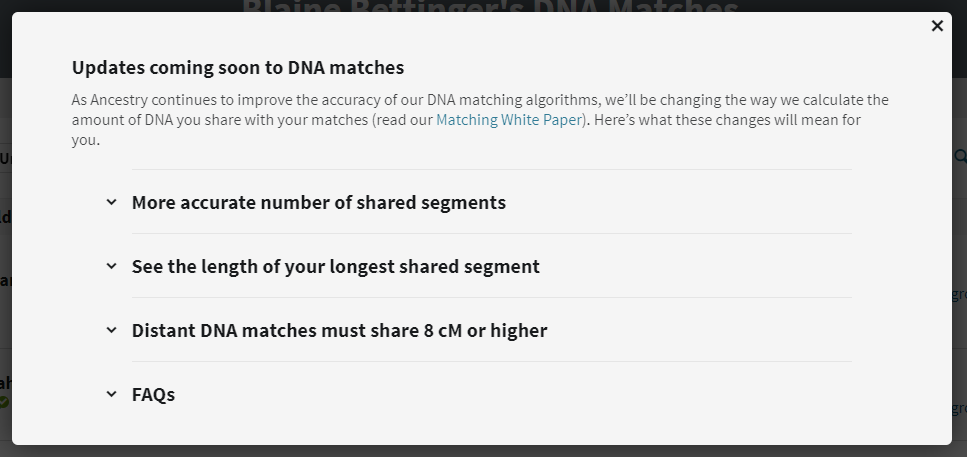
The changes to the DNA match list comprise the following:
- The number of shared segments should improve
From the announcement: “The DNA you share with a match is distributed across segments – short segments, long segments, or some combination of both. Our updated matching algorithm may reduce the estimated number of segments you share with some of your DNA matches. This doesn’t change the estimated total amount of shared DNA (measured in centimorgans/cM) or the predicted relationship to your matches.”
In Genetic Genealogy Tips & Techniques, we’ve noted that the number of shared segments is usually inflated. For example, I share 44 and 49 segments with my parents when in fact it should be exactly 23. It will be interesting to see if the number is 23 after the update. We haven’t found the number of segments to be particularly useful evidence anyway, in most cases.
2. Length of the longest shared segment will be provided
From the announcement: “The length of the longest segment you and a DNA match have in common can help determine if you’re actually related. The longer the segment, the more likely you’re related. Segment length is also the easiest way to evaluate the difference between multiple matches that all show the same estimated relationship. Our updated matching algorithm can show you the length of the longest segment you share with your matches.”
This is a surprising change! Knowing the length of the longest segment shared with a match will be enormously beneficial to test-takers with endogamous ancestry. It will allow them to identify, to a degree, matches that share only small pieces of DNA (and thus much older common ancestry) and matches that share at least one large piece of DNA (and thus more recent common ancestry).
Of course, the efficacy of this improvement depends entirely on the previous improvement, namely the ability to identify the true length of segments shared by a match.
3. Matches that share 6 to 7.9 cM will be eliminated
From the announcement: “Our updated matching algorithm will increase the likelihood you are actually related to your very distant matches. As a result, you’ll no longer see matches (or be matched to people) that share less than 8 cM with you – unless you have added a note about them, added them to a custom group or have messaged them. These changes to the matching algorithm will reduce the total number of DNA matches you have and the number of new matches you will receive. It may also affect the number of ThruLines you may see.”
This is the big one, and the focus of the remainder of this blog post!
There is an updated Matching White Paper available from Ancestry.
The (In)Validity of Small Segments
A very large percentage of small segments are not valid shared DNA, and half of the valid small segments are at least 20 generations old.

As many of you have seen before, I call small segments “poison.” I equate them to poison chocolate-covered candies (because I love non-poisoned chocolate covered candies). If I handed someone a bowl of candies and told them that I had poisoned 50% of them and the poisoned candies looked like every other candy in the bowl, no one would reach in and grab a candy. There are plenty of sources of information about the danger small segments, including “A Small Segment Round-Up.”
Unfortunately, the answer to that scenario is often “Yes, but…” There are no ‘buts’ when it comes to small segments. There is no test or application that can distinguish between false and valid small segments. There is no evidence that mapping small segments, triangulating small segments, or finding small segment matches in shared match groups increases the likelihood that a small segment is valid.
Phasing (separating DNA from a test into the test-taker’s paternal and maternal contribution) reduces false segments, but typically only one person in a match set is phased and the unphased individual may be matching due to a pseudosegment. Phasing also does not resolve the ‘age of the segment’ issue discussed below.
And yes, there are VERY limited scenarios when we CAN identify small segments as being valid, such as during visual phasing (showing recombination at the ends of the chromosomes, for example). Another scenario is where another family member shares a large segment with the match that encompasses the small segment that I share with the match (in which case we’re working with a large segment, not a small segment).
Part of the trouble with small segments is that they are enticing. We can find what looks like great data in the enormous pool of distant matches. For example, it is estimated that as much as 50% of our matches at AncestryDNA are in the 6-7.9 cM range. I have 83,108 matches in the range of 6 to 20 cM. Even conservatively taking 25% of those matches as being in the 6-7.9 cM range means 20,777 matches. And if half of those are based on false segments, that would be about 10,000 matches. How can I not find genealogical connections among those 10,000 matches? Unfortunately, finding the genealogical connection does not mean the segment is valid.
In addition to the validity problem, small segments can be very, very old. The best source of data we currently have on the possible age of segments is from the Speed & Balding paper which is linked and discussed on the Identical by Descent page on the ISOGG Wiki. For example, according to their simulations only 20% of valid 5 megabase shared segments (which is roughly equivalent to 5 cM) are within the past 10 generations. 50% are greater than 20 generations old.
Why is age important? Because when a segment is very old, that means it could have come from a very distant shared ancestor that we may not even know we share. Finding a shared segment and finding a shared ancestor are two separate pieces of evidence that we have to work to combine. That is often impossible to do when working with only small segments, since our trees are all woefully incomplete to the distance from which small segments can come.
But I have a confirmed cousin sharing 6 cM!
Because so many small segments are invalid, and because so many valid segments are so old, the amount of evidence necessary to establish that a small segment shared with a genealogical cousin came from the identified common ancestor is gargantuan.
First, there must be evidence that the small segment is valid. The genealogical connection is not that evidence; that’s circular reasoning (and there’s no science to support that conclusion).
Second, there must be evidence that the small segment came from the identified common ancestor and not from another, potentially unknown and very distant, shared line. This is probably the most challenging of the two issues given the potential age of small segments.
Without evidence for both issues, the genetic conclusion is unsupported.
And see my point above about my hypothetical 10,000 invalid matches. It is impossible for me not to find genealogical cousins among those matches. Believing that I beat the odds is confirmation bias.
But there are valid genealogical connections among those distant matches!
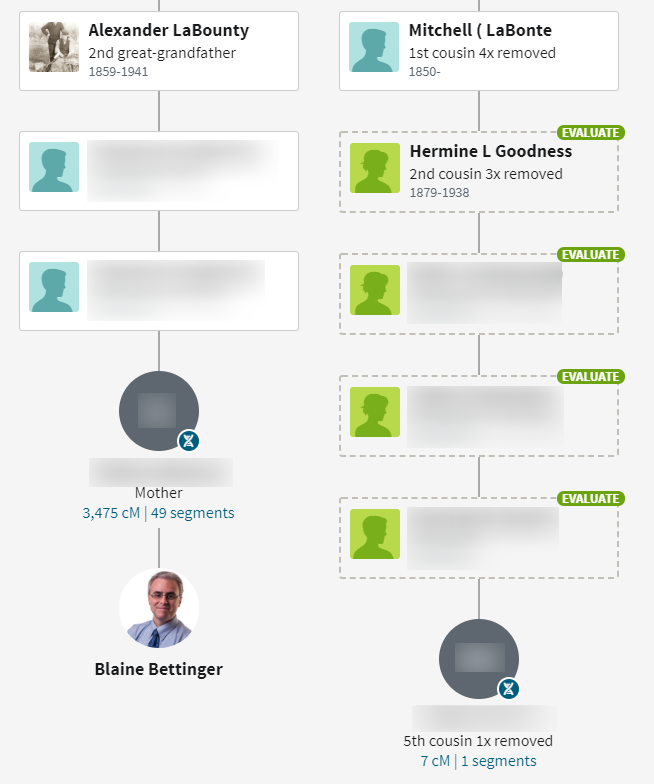
This is absolutely correct. There are very valid genealogical connections among distant matches, including among all the false matches. That the shared DNA is invalid does not invalidate the genealogical connection. This change will result in many genealogical connections being lost. For example, the following ThruLine is based on a small segment and used several trees to find a potential genealogical connection:
I have 69 matches in the range of 6-7 cM that have a Common Ancestor designation, out of 431 total (so 16%).
This information will potentially be lost when the 6-7.9 range is eliminated (although it might still be found among the trees at Ancestry). However, this is for “the greater good.”
Yet another problem with distant matches is that DNA test-takers have no idea that a large percentage of small segments are invalid. Test-takers that know this and use small segments cautiously represent a minuscule portion of the database. That means that most people that see the ThruLine above accept its validity, believe the genetic connection is valid, and conclude that they’ve proven their descent from the ThruLine ancestor. Obviously that’s not the end of the world. But why not prevent these incorrect conclusions when there’s a way to do so?
Losing these genealogical connections is the price we pay to protect current and future test takers from relying on false data. And we have so many other genealogical connections to pursue among our valid DNA matches!
But small segments can identify biogeographical origins!
Small segments have been used to potentially identify very old biogeographical origins. For example, many people with African ancestry have found African matches and this may point to biogeographical origins.
While I hypothesize that it might be possible to point to very high-level biogeographical origins with small segments, it’s important to acknowledge that this hasn’t been demonstrated with any scientific evidence. For example, I haven’t seen an analysis of someone with African ancestry versus someone without African ancestry to see if only one or both can identify African matches. Of course that doesn’t invalidate the approach, it only means we must proceed cautiously.
Indeed, it will be important to consider and study how this might affect people with African or other historically marginalized ancestry. All the available science about small segments indicates that this change will actually improve the method by weeding out false data and preventing incorrect conclusions. Additionally, as more Africans test the number of larger shared segments will increase. Those of us that have been genetic genealogists for a long time have seen several instances of DNA results provided to people of African descent that were later discovered to be incorrect as testing improved and databases grew. Hopefully this change prevents that from happening.
There is a GREAT blog post from Tracing African Roots about the dangers and potential rewards of this approach: “How to find those elusive African DNA matches on Ancestry.”
The road ahead
Regardless of the known issues with matches in the range of 6-7.9 cM, I know that people want to and will continue to use the genealogical data that is currently among some of these matches (such as Common Ancestor hints). You can retain matches in this range by doing one of three four actions:
- Add a note in the match note field;
- Add them to a custom group using the colored dot system;
- Message them; or
- Add a star [this was added by Ancestry on July 18th].
We don’t have an exact date for the match update, only that it will happen in early August.
Blog post round-up
Randy Seaver at Genea-Musings has a round up of other blog posts about the change: “AncestryDNA Changes Coming Soon – What I’m Doing.”
.
In between trying to determine how best to approach saving some of these 6-7cM matches (surname / location etc) I decided to note the number of total DNA matches for the kits I manage and can view. It will be interesting to see how the before and after compare!
I’d love to see the comparisons in the FB group. Maybe a reminder for folks to catch the “before” numbers? 🙂
If there is no test or application that can distinguish between false and valid small segments, a lack of evidence that mapping, triangulating or grouping of small segments helps determine their validity cannot be used to argue that those methods don’t help. You would need an independent method of distinguishing between false and valid small segments in order to evaluate the usefulness of the other methods.
People have done several of these studies. We’ve discussed many of them in GGTT and in various blog posts.
About 90% of my Thru Matches take me down a tree known to me for several generations and terminate in a relative in the 6-10 cM range.
The larger segments are good ones to pursue, although they can be much older than the ThruLine ancestor as well.
Great explanation of why small segments are problematic.
An 8 cM threshold will lose a few genuine matches below 8 cM – but the new threshold will make the match list more reliable for those matches at 8 cM and above.
A little background, if I may. I have been working very seriously on my family tree for just shy of fifty years. I was lucky enough to be raised and live a time in the vicinity where many of my ancestors settled 200 years ago. I had the opportunity to examine many original records like wills and the under-appreciated deed records of my ancestors. As a result of these fortuitous circumstances, I have a pretty complete tree. I know all my 4-greats, and my tree is over 80% completeness out to 5-greats. My records cover several thousand descendants. Virtually all this prior to DNA testing.
But I have found DNA matches to be very useful. It is gratifying to see a new match and know where it fits into the tree, and it helps to confirm all that work (and sometimes point to a new branch). But where to go from here? Of course I am trying to expand the tree further, and to establish more branches. I must delve into more 6c, 7c, 8c, and that means small segments, by the very nature of recombination.
I frankly do not care that It is an old segment, The fact that I share it with someone is a valuable clue. I do not care if we are related in more than one way (usually are at these levels). Again it is just a clue to know of the relatedness.
So, I am distressed to know that in the future, these valuable clues will be unavailable to me. It’s like hearing that the local genealogical library suffered a fire that destroyed some of the books. I would not know that they would have been useful, but I cannot find that out, either.
William, my thoughts are very much in line with yours. The small cM matches have been tremendously helpful to my late 18th and early 19th century research. DNA connections from this far back can’t be anything other than very small. If given a choice between too much information and not enough, I would err on the side of “too much” every time.
Thank you William…my thoughts exactly…just past this week I finally got one family line straightened out by having 2 new relatives with their small family tree…and on was 6cMs and the other 7 cMs…
Because of this new removal…I have to spend my precious time saving up to 40,000 of my 6-7 cMs by marking them…Thank you Ancestry! You’re such a doll!
As usual, a fine summary and commentary! And, I’m especially glad to see that you are still in genealogy. MY problem: I subscribed to DNACentral in March. It appears that since then, there have been no new content. I’ve sent two emails asking if I’m doing something wrong but have not received any reply. Please advise! Thanks!
Thanks for the article; I suppose I fall mostly into the “there are valid genealogical connections among those distant matches” group. I get that there are a multitude of false matches; However – a short story – about a year ago, I discovered the connection with a DNA match that opened up an entire branch of my family tree that was previously unknown to anyone in family (because my GF was orphaned and sent away.) This particular match is a third cousin (and we have no question about this relation after very stringent fact checking!) This third cousin and I share 7 cM of DNA. The discoveries for my family have been immeasurable. In fact, when I look across my matches that I have verified with the paper trail (707 of them as of today), I count 237 of those matches that share 6 or 7 cM of DNA (that’s 1/3 of my “verified” fact-checked matches!) I am very much fact-based in my paper trail investigations; I don’t buy that it was just “luck” that a match of 6 or 7 cM also has a demonstrable shared direct ancestor. While I am glad to have made these connections, it is disappointing and sad to know that they will not occur in the future under the new algorithm.
Thinking more about this, I think it is the statement: “It is impossible for me not to find genealogical cousins among those matches. Believing that I beat the odds is confirmation bias” that gives me some disbelief. Giving thought to some of the astronomical odds:
*Ancestry has 18 millions tests
*they found around 29,600 matches for my DNA
*Assuming ~1/2 are 6-7 cM, that’s ~14,800 results in question
* some much smaller set of those results has “shared matches” with others of higher
match, potentially previously verified to be in a certain family line; just assume 5% or
~740
* I have verified a genealogical connection with 237 of those 6-7 cM matches, selected
based on common DNA matches with other “verified” matches – many of which with
more than 7 cM match, and whose direct ancestor is within the same line as other
shared matches. This seems beyond coincidental.
I agree with the basic statement that you can find genealogical matches in that set. But to find matches in the same genetic line, as often indicated by “shared matches” with other results >7 cM (sometimes 20-30 cM) cannot have been a lucky coincidence for me 237 times (I should play the lottery if I am that lucky!) There are many more matches that I have researched and not been able to find that common direct ancestor (at all levels of matching DNA). If I randomly choose one of those 6-7 cM connections with no shared matches, perhaps I could eventually find some link, but in most cases, not. However, to find that connection in the same line as a shared match, in a small segment match specifically selected due to those common matches – not once or twice, but 237 times… even with all of the false matches that are possible and even with the possibility that the DNA cannot be verified to be from a particular ancestor (and thus is considered “invalid”) it appears that I was able to pick the needle from the haystack, again and again. I have to conclude that those 6-7 cM matches offered some significant genealogical value. I think Ancestry’s position lowers the bar, and along with their ongoing resistance to adding chromosome browser, shows that they have set their course to oversimplify and be a low-quality provider of DNA tests to the masses.
The Ancestry spin that removing DNA matches under 8cM Is a benefit to me is absurd. How will removing potential research data be a benefit, even if the “success rate” of confirming the match is low? The fact that you can find successful confirmation of matches under 8cM is enough reason to retain them in the database. This change is to benefit Ancestry, not me. Whether it is to free up their server space or to create a “premium” DNA access level or whatever benefit they are trying to accomplish, I would much prefer to be told that than to be subjected to the disingenuous “we’re doing this for you”.
I don’t think you read the blog post? You can’t confirm DNA under 8 cM.
So, that is why I was never able to TG using 8 cMs…………..
I don’t understand that statement.
I have many a match connected who are under 8cM and they are all connected by census records, birth records, death records and marriage records. If that isn’t ‘confirming’ then what is?
I would understand it, if it were attributed to a country/society that held no long term ‘official’ records, like say, Somalia, Afghanistan, etc etc.
For the record, my lowest cM match confirmed by a ‘paper’ trail and with multiple shared matches of DNA tested individuals and their associated paper trail, was 4 cM. Granted, it required a tree of over 15K people and governments that kept accurate records back to the 1700’s- In this case England and Scotland. A tree that size would eliminate the majority of tree builders, certainly within Ancestry.
To claim that “You can’t confirm DNA under 8 cM.” becomes more and more ridiculous, as the years progress as subsequent generations take DNA tests, and data collection/storage allows more paper ‘trails’ to be kept.
For instance, if I am a match with an individual (distant relative) with, say, 100 cm, all is good, but if my grand child was to take a DNA test and the other individuals’ grandchild were to take a DNA test, the cM’s between them (the two G-children) would be well under 8cM, and what you are saying is that the link between them would be impossible to prove… Do you see how ridiculous that statement of yours is?
How have you proven that the segment is valid? How does finding a common ancestor prove that a tiny segment is valid?
If I take a random person off the street and say find a common ancestor, with sufficient trees everyone would eventually find a common ancestor. If you lower the threshold to 4 cM at GEDmatch, good luck finding anyone you DON’T share a 4 cM segment with.
Your unwarranted anger is misplaced. Direct it to trying to improve DNA testing so that microarrays don’t create pseudosegments
When there’s only one segment shared, the chances are higher that it’s an “old” segment (correct me if I’m wrong). So the improved estimate of number of shared segments may provide some pretty useful information.
One thing that seems disappointing, and one of the reasons I’m more in the “they’re doing it to save money, not to help me” camp is I don’t think there’s anything magic about 8. Maybe next year, it’s 9. If there’s no way to confirm DNA under 8 cM, I think there’s no way to confirm DNA under 9 cM, etc. I have some distant matches that are a significant number of cM and only one segment. I am developing good theories on where those segments came from, due in part to some lesser-cM matches. I’m not necessarily looking for those distant ancestors – I’m looking for the more recent one that this segment came through, and it’s been clouded by NPEs. I don’t really care if a lot of them are false. Initially I care more about casting a wide net. Having more matches increases the odds I can follow this clue further, with cross-references to other testing and matching sites and tools.
Thanks for the article.
Although I don’t believe the “for the money” argument makes any sense, the rest of your comment is one of the most astute I’ve seen all day. There’s absolutely nothing magic about 8 cM. Plenty of 8 and 9 and 10 cM segments are false. Personally, I think they should have gone right to 10 cM, where they could reasonably stay for a very long time.
Thanks.
“For the money” refers to less matching. This means smaller databases, storage space, AWS servers, etc. These things cost actual money, and most likely, the cost does not scale linearly. It’s not my original idea, but it does make sense to me. I used to work in the database software industry, and I draw on that experience in making this claim, though I don’t know anything about their infrastructure specifically.
If it didn’t cost them any more, and they were doing it for the user, they could make it a default option. There is already a filter option for cM. Even if doing it for the money, it may be a reasonable trade-off, but some long-time users understandably feel the loss and don’t generally think they have any input or control, especially when they understand the caveats and have invested time in something that may now simply vanish. So it’s hard for me to see any argument that they’re doing it for me.
It’s now taking me some time to see if I can find all of the ones I have researched and have in my research files (across more than one Ancestry kit) to try to get them tagged in hopes they won’t go away before I’m done looking at them. There may not even be very many of them I care about, but it takes me time to go through them to determine that. It seems like a loss mostly because it’s time I didn’t know I’d have to spend.
Turning it off but providing an option to turn it back on seems like a good compromise.
Even at 1% chance that a match is valid, that’s still better than picking someone from the population to see if they might be related. Especially if they have a name you’re looking for, or are a confirmed match to someone else you’ve confirmed a relation to. I think that’s what people are unhappy about, losing that hint. The people who don’t do proper research to confirm connections aren’t the real target audience, as far as genealogists are concerned.
Great! Now I’m craving M & Ms! 🙂
Haha, I know the feeling!
So, then for our self preservation, we better also be marking our 8,9 & 10’s right now and as they come in the future because we could be in peril of losing them too with little advance notice.
Good article! It went a considerable way in allaying my fears about losing these potential matches. However, I don’t think you have shown that the small segments are not potentially useful. Even if we can’t tell which matches are valid, and only a small percent are, they could still be a richer ore to search for, say, relatives within 10 generations.
As for cost considerations, that actually seems like a potentially valid reason for them to do it. As more people test, the amount of server and time resources to make calculations will grow, probably much faster than linearly. On the other hand, they could probably save almost as much simply by making the default be 8 cM (both for totals and comparisons), and allowing comparisons to a smaller cutoff for those few people who might want to do that.
Right at the time of the cease-and-desist for 3rd party tools, I did one last DNAGedcom download of all the kits I manage, so I can compute how much of my matches are <8 cM. 58% of my roughly 31,000 matches fall in that category. Ugh! My problem now is I need to clean up my notes from 2014 and 2015 when I used to write "no shared matches" or "doesn't match mom or dad". I especially don't want those to stick around! I'm looking forward to seeing segments shared w/ my dad (55 segments) and my mom (77 segments) when all is said and done.
If one shares 7 cMs with an ancestral grandparent somewhere back in time with another person and each person shares the grandfather, but each comes from a different wife, that skews the cMs.
Because if each person was descended from the husband and wife couple, instead, would they have shared abt 14 cMs? Just hypothetical……
Thank you, Blaine!
I’ve been hunting 5th-8th cousins for a couple of years hoping to find another tree with names I recognize. I have 21+ DNA matches to another branch of the same surname. I haven’t gotten any farther back than before. I do now have a suspicion of a common ancestor but it’s still iffy. I know I am grasping at straws. I know the matches I’m getting could be through a different person than the one I’m targeting but all I want to see is their line back to New Jersey. No one can go back to 1776-1784. So I keep chipping away at those 5th to 8th cousins hoping at least one of them shares my suspicion.
Blaine,
A couple of specific DNA cM ‘purge’ questions…
Could you verify that the purge of matches is only for those less than 8 cM?
And, importantly, is that amount the amount of total cM which we currently see shown for each match, or will it be for the largest segment amount? In other words, might we also lose matches who currently show as having total cM amount above the coming new cut-off amount because none of the segments is 8 cM or more?
The other cM question:
Because they not so long ago started rounding and showing no decimal amounts (matches which earlier showed, for example, 7.7 or 7.9 cM are now shown as 8 cM), it seems that we need to also check those 8 cM matches (in case they were actually less than 8.0 cM decimally) if we really want to protect/preserve them… 8 cM may actually be 7.6, 7.7, 7.8, 7.9 and get purged? Is that correct?
Regardless of how valid the lower cM is in proving descent from a specific shared ancestor, I have found value in many of these matches in furthering genealogical research and getting leads — both genetically and genealogically — and would like to preserve many of them on the site in association (like ThruLines, etc.) with the ancestors I’m researching. I’m working primarily with 4-6th great-grandparents where, according to your cM Project on DNA Painter, it is not unusual to share these lower amounts with cousins in this range.
I ask the specific cM questions because I have over 117,000 matches on Ancestry.com, and all but about 4,000 are in the “Distant matches” range (6-20 cM) … over 113,000. And, a kit of an elderly ‘brick wall’ relative who is a generation closer to that wall, which I use to research on those lines, has over 166,000 matches in that Distant range. Not to mention several other related kits also with more matches than I.
So you can see I need to prioritize very precisely to be able to save as many of the most meaningful (to me) ones. I imagine there are others out there who value your knowledge and expertise and might be wondering the same thing…
Could you please clarify the cM questions?
And, of course, any other advice to make this task easier or faster, is most welcome.
Thank you for your guidance and helping us all learn and understand!
This article was news to me and I was not aware that Ancestry plans to cut a couple of levels of cams. I am not surprised, however, as lately the slowness of Ancestry trees has been unbearable. Even adding new people has taken retry after retry and still unsuccessful. Am I the only one? Is it my computer?
Is Ancestry not under control of an LBO? If so, it is a given that additional investment in servers is not in their future. Perhaps anew owner with a capital infusion would solve many ongoing problems.
I am sorry Ancestry is doing this. I use the matches to point me to possible branches to pursue in written records. My hobby for 20 years been using Italian civil records 1809-1900 , US written records, and correspondence to put together one big tree of the residents of a small Italian town and follow immigrants from that town. DNA matches, even the 6 and 7 ones, especially if the tester has even a small tree or a recognized surname, can point to a new branch or find lost ones.
I have used “6” matches to find at least two direct ancestors who “disappeared” but with the lead of a couple of 6 matches, I tracked my lost ancestors thru a records search online at the Italian records site Antenati. Without the DNA hints provided by these “6” matches, I would never have thought to search thru the records of the other Italian towns for my lost grt x3 grandfathers. I know only barebones DNA science. What I do know is is old fashioned using of hints, hunches, and painstaking records searches. Only a small portion of my 6 and 7 matches raised my interest, but they also provided the only clues for me to hunt certain branches. What a loss for researchers like me when they go.
Thank you, Blaine, for the detailed explanations and thanks to the several reply people for their comments. My own “rule” about low scoring matches has been to look hard at their shared matches. If they have several shared matches with really solid higher scoring matches that has been a clue that the low scoring match should be taken seriously. Jonathan Brecher’s cluster tool has been useful in identifying the many low scoring matches that have these shared matches. Thus, a fair number of my 6cm and 7cm matches have been identified and fit nicely into useful clusters, adding data for specific family branches. In round numbers, the Brecher tool identified about 600 matches for me in the 6-7cM range that I have found to be useful — and likely correct matches. It seems a shame to lose these extra matches both now and in the future.
Thank, Blaine. I’ll certainly take the <8cm change if it means we can now see the largest segment size of our matches. I do hope this change is indicative of more changes to come at AncestryDNA. Chromosome browser here we come! 🙂
I agree with you totally on the lack of value for small cM matches while doing genetic analysis. What I worry about is the effect of removing the small matches on the behavior of subscribers to Ancestry. I am an old psychologist who studied BF Skinner’s work on schedules of reinforcement. My use of AncestryDNA can be viewed like a rat pressing a bar for a food pellet. Each time I look at my matches on the web site, I may be reinforced by finding a new match. Some have high incentive values and some have low incentive values, but each provides me with reinforcement because I have covid-induced time to fill and I enjoy the hunt. Skinnerians would call the reinforcement schedule here a “variable interval schedule”: Reinforcements are made available at unpredictable intervals and the schedule is independent of my behavior, but I only receive the reinforcement with a response. Reinforcements on a Variable Interval schedule are very good at maintaining steady responding over long time periods. For Ancestry this means that users are likely to distribute their web activity fairly evenly and they are likely to keep subscribing. If reinforcements stop, then the responses will maintain for a while but then become sporadic and eventually Extinguish. If the interval between reinforcements is too long, responding is not maintained consistently because Extinction will occur between reinforcements. So, to my point. The biggest effect of eliminating the low cM values may need consideration, not just in terms of their value in genetic analysis of the tree, but also in terms of the effect on the behavior of Ancestry’s user base. In the last 3 wks, I have received 136 ‘reinforcements’ for looking at my match list. Eliminating the 6-8cM matches would cancel 81 of those. From the perspective of a dedicated and sophisticated genetic genealogist, that may seem like a good thing. From the perspective of a company that relies on consumer behavior, that may not be a good thing. Time will tell.
And, BTW, I have found over 300 matches with 6-7 cMs AND a tree match. The feature Ancestry introduced last year (the ability to search upstream in the tree of your match) has been invaluable to me.
David Tieman you had me giggling at this – “My use of AncestryDNA can be viewed like a rat pressing a bar for a food pellet. Each time I look at my matches on the web site, I may be reinforced by finding a new match.”
Hilarious! I too am like a rat, being rewarded with new matches.
I have marked many of my smaller cm matches, to keep them.
I think 5 cM is not the same as 6 cM is not the same as 7 is not the same as 8 cM. I would bet they decided they needed to do this because of “backend systems overtaxed.” That’s reasonable I suppose. No one wants a website that can’t function. But if the cut-off point were based on specific desirable positive predictive value, I doubt it would be a convenient round number. The number of generations to choose is arbitrary, but I would think they could choose a cut-off based on something more along the lines of “this gives us a 75% positive predicitive value for the match being true at 7 generations or less”…something like that.
Hi Blaine, Regarding your comment on the inflated number of matching dna segments, you mention that the number of segment matches you have with your two parents are in the 40s when it should be 23, each segment being an entire chromosome. Is it possible that in the case of parent child matches each cross over event within a chromosome increases the number of segments that the parent matches with their child? I looked on gedmatch at a women and the matches she had with two of her daughters and observed that each chromosome was broken into 2 – 4 segments where they matched. Just a thought. I have no idea if this makes any sense.
Ancestry has made a marketing error. IF they intended to purge small 6-8cM segments, and offered a decent explanation plus a one-time simple way to keep those segments, only a small fraction of subscribers would be interested. Who is really going to pursue thousands of leads now, plus more in the future? Instead, the it’s-going-to-happen approach has put those same subscribers in a negative state of mind about Ancestry’s intentions, and those subscribers happen to be the more serious genealogists and long-term customers. BTW, I do use those small segments, but as leads to family trees that extend past mine.
I would like to know what a good (or reasonably, bang for the buck, worth the effort) lower threshold for cMs is. My mother has (had) thousands of matches, but only 216 (and this is on Ancestry) that are 20+ cMs. (And I’ve not yet confirmed a single match genealogically. Though I keep trying!)
I assumed if I found a cluster of lower cM matches (they match with each other, their chromosome segements overlap (GedMatch/FTDNA/DNA Painter), common surnames found, etc;) then I could comfortably undertake the task of trying to triangulate (1) *their* common ancestor; and (2) *our* connection to them via the common ancestor.
An example, I just put together a cluster of folks, that range from 25 cM (without a tree) down to 6.4 cM (with a tree, which allowed me to connect them to someone else at 8.9, who lived in the same district as some same surnamed folks in 14-15 cM range, one of which match the 25 cM person.)
If I can work them into a cluster, aren’t the odds are better, despite the low cM, there is a match?
This is what I need. Thanks you
I was lucky enough to have time to save very one of the 6 and 7 Cm DNA MATCHes for the 6 accounts I manage. My wife and I spent over 200 man hours doing this. Since then, I have had Ancestry automatically make Common Ancestor matches for 16 of these 6 and 7 CM. Was well worth the time.
While small segments can be helpful for identifying genealogical connections, they cannot be utilized as genetic evidence to support those genealogical connections, unfortunately. At least based on all available science.
To claim that “You can’t confirm DNA under 8 cM.” becomes more and more ridiculous, as the years progress as subsequent generations take DNA tests, and data collection/storage allows more paper ‘trails’ to be kept.
For instance, if I am a match with an individual (distant relative) with, say, 100 cm, and there is a proven, sound paper trail, all is good, but if my grand child was to take a DNA test and the other individuals’ grandchild were to take a DNA test, the cM’s between them (the two G-children) would be well under 8cM, and what you are saying is that the link between them would be impossible to prove…
Do you see how ridiculous that statement of yours is?
Great blog subject. As I do more research on my unknown grandfather – a man probably born around 1860-70 I grieve the loss of these under 8 cM matches. My father was quite old when I was born. He was also born and raised in another country. Most of the people alive today taking DNA tests would be 2-4 generations removed from my grandfather. The snatches of DNA are a critical clue ir a door opener – of course they don’t replace the necessary research, interviews, testing, paper documentation but the loss to this research is irreplaceable.
Independently of my grandfather saga, I regularly find a helpful match I did save when I did become aware of the DNA purge. Unfortunately I then have to “prove” the match through emailing a screenshot. It casts an attitude of mistrust over a request to view a tree or questions about a shared ancestor.
I saved about 20,000 thru stars, dots and notes. I lost about 20,000. I saved by nationality the most likely matches associated with my paternal grandfather. I had been unable to how far flung his relationship with my father’s nation.
When I came across this article, I thought it important to post my own post mortem to this Ancestry corporate business decision in the hopes that it may trigger more conscientious choices in the future.
I appreciate the quality of your blog and the wonderful, well informed feedback readers give in return. Thank you to all.