My grandmother Jane died in 1984 when I was just 8 years old. I have some really great memories of her, faded with time but still filled with emotion. Bath times, spending time with her in the summer, newspaper hats, chrysanthemums.
However, in addition to those memories, she gave me a very unique genetic heritage. She was from a region of the world with a high degree of admixture, and thus it is from her that I obtained my Native American mtDNA, my Native American, African American, and Spanish autosomal DNA. It is an incredibly rich and fascinating genetic legacy.
In an attempt to learn more about my grandmother’s genetic heritage, I’m using GEDmatch’s new Lazarus tool to try to recreate as much of her genome as possible. Join me on the journey, and learn about this new tool.
Lazarus
Lazarus is a relatively new Tier 1 tool at GEDmatch that allows users to create what are called “pseudo-DNA kits.” There is a help page called “Lazarus” with a description and examples, for more information. From the description at GEDmatch:
Generate ‘pseudo-DNA kits’ based on segments in common with your matches. These ‘pseudo-DNA kits’ can then be used as a surrogate for a common ancestor in other tests on this site. Segments are included for every combination where a match occurs between a kit in group1 and group2. The resulting are combined to create the final kit. It may be useful to save copies of the next page results, for future reference.
Therefore, Lazarus creates a pseudo-DNA kit by identifying DNA shared between the people you put in GROUP 1 and the people you put in GROUP 2:
- GROUP 1 must be DESCENDANTS ONLY of the target Lazarus person – children or grandchildren, up to 10 of them. Note that if you have a child, there is no reason to include children of the child; you shouldn’t capture any additional DNA by adding them (and at least one person has suggested that adding the child decreases the quality/effectiveness of the Lazarus, but I can’t verify that myself). Do NOT put cousins, siblings, or parents in Group 1!
- GROUP 2 must be NON-DESCENDANT RELATIVES ONLY of the target Lazarus person – siblings and parents are preferred. For most of us, it will be very challenging to recreate most of the target Lazarus person’s genome without one or more siblings and/or parents. However, you can still recreate much of the genome with more distant cousins, as we’ll see below. Do NOT put children, grandchildren, or great-grandchildren in Group 2!
- SPO– USE should be the spouse of the target Lazarus person. This is an optional field, but note that the spouse MUST be the parent of ALL kits listed in Group 1. I’m not entirely certain how the SPO– USE entry is utilized by Lazarus; I’ll try to get more information for that.
Think of Lazarus as a very simple and largely blind algorithm – all it does is compare the DNA in Group 1 to the DNA in Group 2, and then combines all the matching DNA into a single kit. It doesn’t know what you’re trying to do, it doesn’t figure out the relationships between the people in Group 1 and Group 2, and it will not (and cannot) tell you if you made a mistake. So you have to do all the thinking for it.
The most important rule is to NEVER EVER PUT A DESCENDANT IN GROUP 2!! The second-most important rule is to NEVER EVER PUT THE SAME PERSON IN GROUP 1 AND GROUP 2. If the goal of Lazarus is simply to compare the DNA in Group 1 to the DNA in Group 2, and you put the same person in both, you’ve essentially re-created the genome of that person. For example, by putting just myself in Group 1 and Group 2, the Lazarus kit comprises an entire genome, 3781 cMs. It works because it doesn’t analyze what I put there, so be careful and read below about ways to check the Lazarus kit for errors!
EXAMPLE
Alright, let’s say the target Lazarus person is DAD, and you have DNA kits at GEDmatch for the following people:
- Yourself
- Your Mom
- Your full sister
- Your full brother
- Your Dad’s brother
- Your Dad’s sister
- Your Dad’s mother
- Your Dad’s second cousin
- Your Dad’s fourth cousin
Can you cluster those into GROUP 1 and GROUP 2? GROUP 1 will contain: Yourself, your full sister, and your full brother. GROUP 2 will contain: Your Dad’s brother, sister, and mother, as well as his second and fourth cousins. And your Mom can go in the SPO– USE field.
The Results
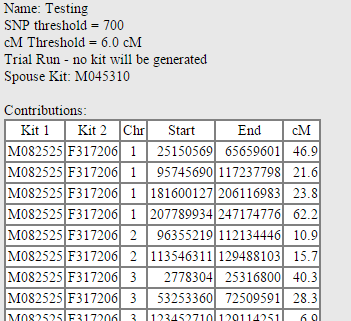
When a kit is processed, the results page will have several sections, starting with a first section showing every matching segment between the people in GROUP 1 and the people in GROUP 2. In the example below, I use myself (GROUP 1), my paternal aunt (GROUP 2), and my mother (SPO– USE) to recreate my father’s genome (who is living, but this is as an experiment). The “Contributions” are every match between me and my paternal aunt:
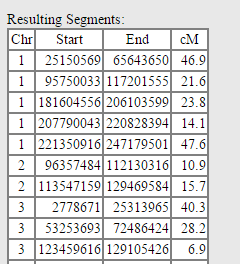
And the next section shows an organized compilation of all the segments used for the new Lazarus kit:
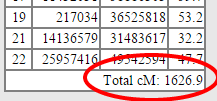
At the end of this column is the most important number on the page:
BE SURE TO RECORD THIS NUMBER!!! IT IS VERY DIFFICULT TO GET IT LATER!! IN FACT, SAVE THE WHOLE PAGE!!
That number, circled in red, tells you the size of the final Lazarus kit. The closer that is to about 3,700 cM (a full genome), the better. A created kit will be eligible for batch processing (and one-to-many matching) ONLY IF the total cM is 1500 or greater. Smaller kits will only be used for one-to-one matches.
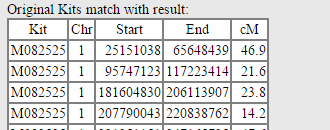
The final column shows the original kits with the utilized segments:
If you’re below the 1500 cM threshold, the kit should be ready almost immediately, or soon thereafter, for One-to-One comparisons. If you’re at or above the 1500 cM threshold, you’ll have to wait a bit for the kit to tokenize, and then it will be ready for One-to-Many comparisons.
So that’s how to use Lazarus!
Checking for Errors
Once you have a kit, how can you check to make sure that you haven’t made any serious errors?
First, for your own satisfaction use the “DNA File Diagnostic Utility” tool at GEDmatch to make sure that everything is as it should be. If you were above the 1500 cM threshold, does the Diagnostic indicate that it has been processed for One-to-Many? (You will have to give this a few days for the kit to be processed, of course).
Second, what was the size of the Lazarus kit you created? If you used one child in Group 1 and your Lazarus kit is 3700 cM, you did something very, very wrong. A child will only have half of his parent’s DNA, and that half can only be used for the Lazarus kit if the people in Group 2 are close relatives (like aunt/uncles of the sibling), and if there are several of them. If you have one child and one grandchild in Group 1 and your Lazarus kit is 3700 cM, you did something very, very wrong. You will have to use common sense to estimate how big the Lazarus kit will be. You are limited not only by the amount of the target Lazarus person’s genome not only passed onto Group 1, but also by the amount of the target Lazarus person’s genome shared by those in Group 2, AND by the amount of overlap between the two groups. That’s a total of three very important limitations.
Also, use One-to-One Comparisons with the Lazarus kit and one or two kits used in Group 1 and Group 2. Does the amount of DNA shared between the Lazarus kit and the person you used in Group 1 make sense, or is it too much?
Lastly, use One-to-One Comparisons for the Lazarus kit and the other side of the family! For example, if you rebuilt your father’s genome using Lazarus, do a one-to-one with your Mom and the Lazarus kit. If it matches (and your parents weren’t related), you did something wrong. Or try the Lazarus kit and a maternal aunt, cousin, or other maternal relative. If there is any sharing (and there’s no known relationship), then there’s a problem. Now, of course, this may not work for an endogamous population (in fact, Lazarus will be tricky – but not impossible – for endogamous populations). As noted on the Lazarus page of the GEDmatch wiki: “If you find any matching segments, look at the ‘Contributions’ list on the results page to see what kit(s) the matching segment(s) came from. Remove that kit from your entry form, and generate new results.”
Next Time
In the next post, we’ll identify kits for my grandmother’s Lazarus project!
.

Thank you! I’ve been wondering how to best use this tool. Looking forward to your insights as you go through the process.
Thank you Nan! Best of luck with your own Lazarus project!
I am trying to reconstruct my Dad’s Genome. There is Myself and my Sister, my Mom (so not related to him) and his has numerous 1st Cousins. Any thoughts on how many CentiMorgans we can get for him?
Nice article on a subject that I’ve been very curious about (the Lazarus tool). I’m looking forward to your next post on this topic. I have a question regarding your example, though. I can understand the benefit of using the Lazarus tool when the situation is not the recreation of a pseudo kit of one of the parents when both a child and the other parent have tested. As in your example, however, if the goal is to generate a pseudo kit of dad’s results, and you have the test results of both a child and the mother, why wouldn’t you just use the GEDmatch phasing tool?
It would be very interesting to directly compare the pseudo kit of dad’s results generated by the phasing tool with the pseudo kit of dad’s results generated by the Lazarus tool to see which is more accurate. The accuracy of both the phasing tool and the Lazarus tool could be even further evaluated by comparing the pseudo kits generated for dad by both tools to dad’s actual results using a case where one child and both parents have tested.
Dan – phasing using the three children’s kits – which would each have to be done individual – would only recover the 50% each of them independently inherited from their father. The three of them would have most of their father’s DNA, but it wouldn’t be combined into a single file.
Theoretically, and this would require experimentation, the three children alone could be used in GROUP 1 and in GROUP 2 to combine them into a single Lazarus kit.
In rereading your post, perhaps my questions are a bit premature (seeing that your father is living and this is an experiment), and that the type of evaluation I suggested is exactly what you have planned for the next installment. My apologies if I’m jumping the gun.
Finally reading this. Note that you cannot use the spouse kit unless it’s real. That is, if I created a kit for my father using Lazerus, I cannot use that as a spouse kit for my mother’s Lazerus.
Thank you Israel. Do you know if that is because the GEDmatch algorithm detects the spouse as a Lazarus kit and as a result declines to utilize it? Or is that because the Lazarus kit is likely not a complete DNA profile?
Does this algorithm use base pairs or the resulting segments of the groups to derive the DNA profile?
If the algorithm uses base pairs, is there some place where one can we what the algorithm is actually doing? How it does the comparisons and how centimorgans are calculated for each segment?
Thank you
If you are going for finest contents like me, simply visit this site daily as it gives feature contents, thanks
So, don’t have the Spouse’s DNA, but I of have a niece of the spouse. Would there be value in putting the niece’s kit in?
My grandfather is the son of two first cousins. His maternal grandfather and paternal grandmother were brother and sister. He married the granddaughter of his {maternal grandfather and paternal grandmother’s} double first cousin. Is that too tanged up for Lazarus? Thank you.
GREAT article! Glad I found it when I did and didn’t waste anymore time not understanding why I wasn’t getting results. I’m was trying to recreate DNA for my grandfather. He was adopted in 1905, no known siblings, biological parents unknown. I had samples for category 1 but was trying to use “Cousin” samples for category 2 but they were pretty far removed.
Have you written any articles on any of the other GEDmatch tools? Any suggestions or articles for using DNA to uncover old adoptions?
I tried to create my father’s Lazarus kit using my mother in Spouse and my bothers and me in group one. It did not create results. Must there be non-descendants in group 2 also, or can we do a “super-phasing” using this. Is there any other way to capture what must be about 70% of his DNA the three brothers share.
A good tip to use when thinking about downloading music
is to make sure you aren’t tying up your internet connection with other things.
Constant dialogue through these platforms is bound to not just positively influence your popularity but also generate greater sales.
My brother was a drummer and my sister was a pianist and a singer.
I have noticed you don’t monetize your blog, don’t waste your traffic, you
can earn extra cash every month because you’ve got high quality content.
If you want to know how to make extra bucks,
search for: Mrdalekjd methods for $$$
If you have the Family Finder results for 2 Children and Soon for their Mother. How many CentiMorgans could you get to?
Hey Blaine, appreciate the strategy used as it has several numbers of options to find the exact match. Check out these other websites that use a different way of finding the match by taking a DNA test here. Take a look at it here. https://www.dnatestreview.org/