What is BigY?
BigY is a Y-DNA test offered by Family Tree DNA. For a limited time only (August 2017), Family Tree DNA is offering BigY to existing customers for $395 (down from $575).
Typically, most Y-DNA tests taken by genetic genealogists are either STR or SNP tests. In contrast, BigY is a Y-DNA sequencing test available to male test-takers. The test sequences approximately 12 million base pairs of the Y chromosome, and identifies SNP results within those 12 million base pairs.
From the BigY FAQ (also see the 2014 BigY White Paper):
The Big Y product uses next-generation sequencing to reveal genetic variations across the Y chromosome:
- Targeted Non-recombining Y-DNA sequencing.
- Illumina HiSeq 200.
- 55X to 80X average coverage.
- Around 11.5 to 12.5 million base-pairs of reliably mapped positions of non-recombining Y chromosome.
- Analyzed using Arpeggi genome analysis technology for improved variant calls.
All samples are processed in-house using our custom laboratory methods and informatics. Your sample never leaves our company and is never shared with outside vendors..
The raw data from the test, VCF and BED files, are available for download on the results page.
What do the BigY Results Look Like?
As with all other tests you take at Family Tree DNA, your BigY results will appear in your results dashboard:
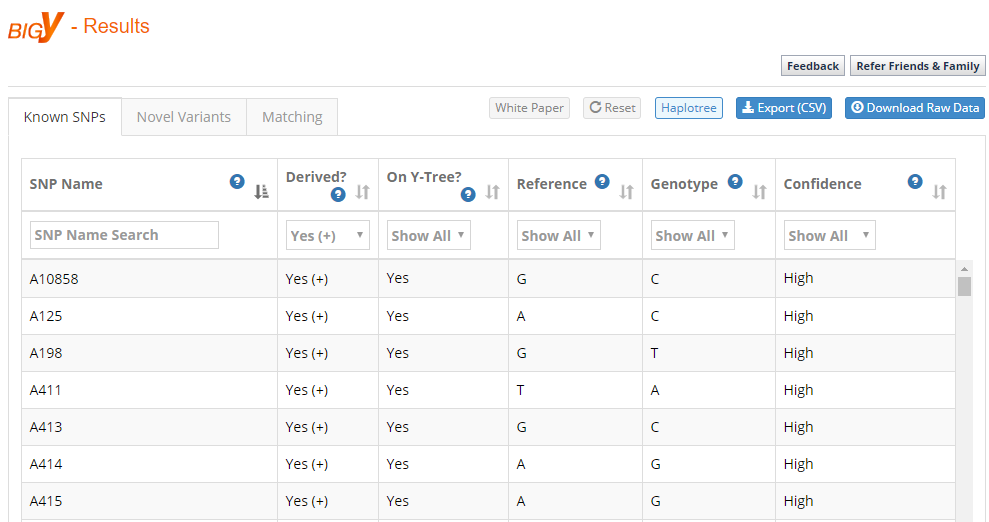
Clicking on the BigY Results tab shows you the list of SNPs from the test based on one or more filters. For example, as shown in the figure below, you can filter based on whether you are ancestral (-) or derived (+) for a SNP, whether it is on the Y-Tree, and other filters.
For example, if I filter by derived, I have a total of 645 results. This means that I am positive for these SNPs, that I have a mutation at these SNPs and no longer have the ancestral value. Notably, these 645 SNPs are out of a total of 36,372 SNPs.
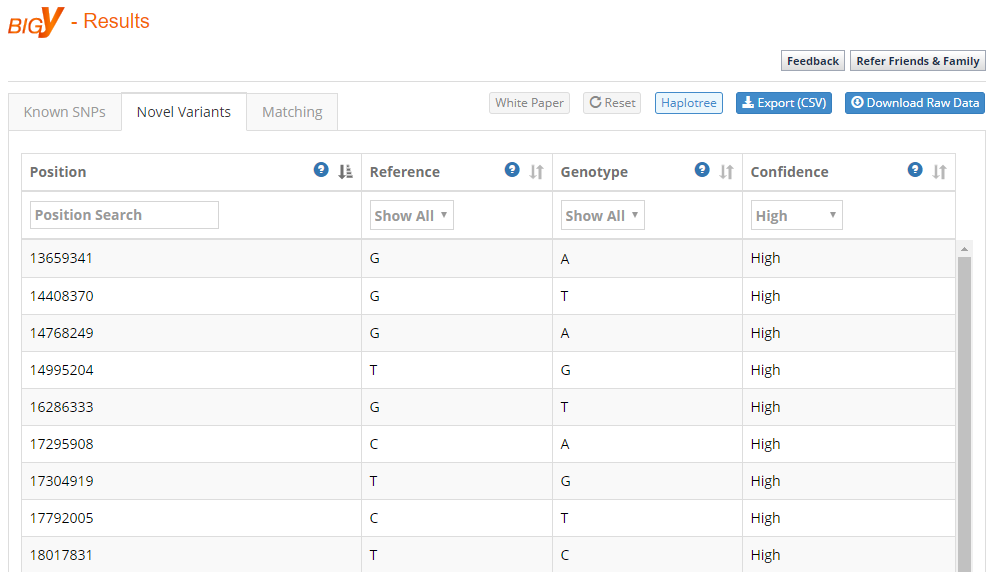
I can also see my “Novel Variants,” as shown in the figure below. Now, this does NOT mean that I am the only person that has this result at this SNP location. There could be thousands of people that have these results, or there could be just me. It simply means that it has not yet been placed on the Y-Tree. Figuring out who has these SNPs, and how they fit onto the Y-Tree, is one of the goals of BigY sequencing. Indeed, we have learned an incredible amount about the human Y-DNA tree as a result of BigY testing.
Eventually, all of these “Novel Variants” will hopefully be mapped to the Y-Tree and I’ll perhaps only have a couple of results that are truly novel to me or a small group of people (presumably closely-related individuals).
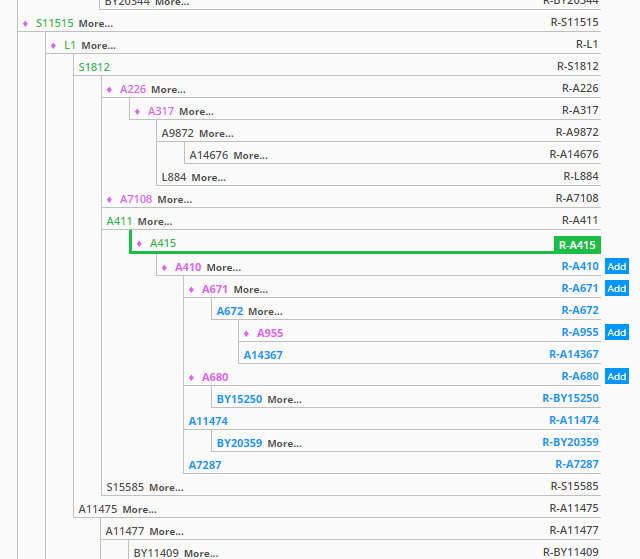
I can also click on “Haplotree” to get a quick drop-down of my spot on the human Y-DNA tree. As you can see, my terminal SNP is R-A415. Before BigY, my terminal SNP (based on previous SNP testing) was R-L1. So you can see that BigY helped push me further out on the Y-DNA tree (in fact, BigY helped create these additional twigs of the tree underneath L1):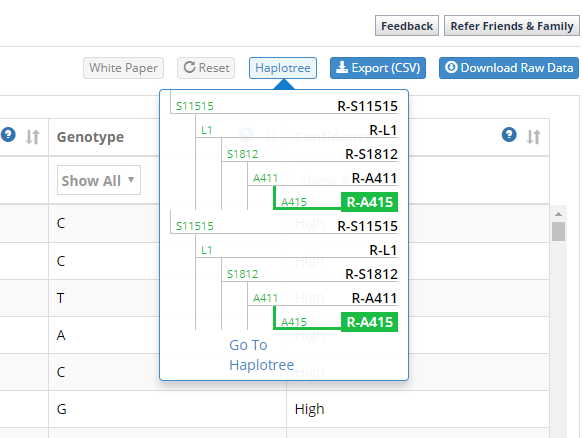
Clicking on “Go to Haplotree” expands the tree and shows a little more data:
Shortly after the BigY results were first available, Family Tree DNA added a matching option to the tool. This compares your results only to other BigY test-takers, and matches appear to be ranked based on: (1) the number of Shared Novel Variants; and (2) the number of total Matching SNPs.
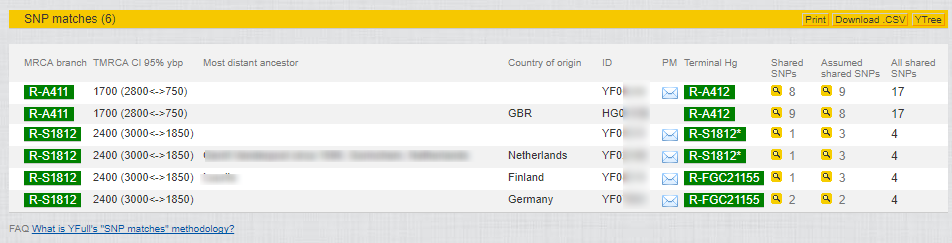
You can see in my match list (this only shows a few of the total 211 matches) that I have a couple of matches with no known SNP difference, who share 8 Novel Variants with me (and almost 30,000 SNPs matching). 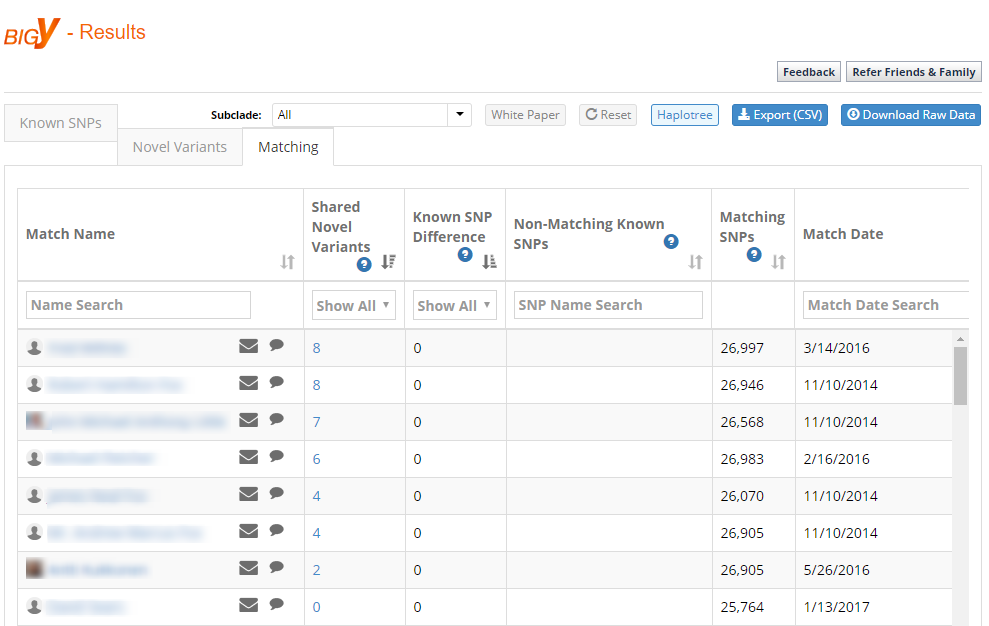
However, finding random matches can be very misleading. For example, the third match down with whom I share 7 novel SNPs has tested his STRs out to Y-111. When I compare his Y-111 results to my Y-111 results, I discover a Genetic Distance of 24 (and a GD=10 at 67 markers). Thus, he is not a close paternal relative, at least not on a timescale where we’ll find any records. But we are almost certainly related within the past 1,500 years or so, and together with other similar matches we will likely learn a great deal more about our particular branch of the Y-DNA tree.
(Discovering this took a convoluted path, but I’ll share it in case you find it useful. First, I found him as a GD=0 perfect match at 12 markers where he listed his terminal SNP and his most distant ancestor. Second, I went to his surname project and found his Kit number. Third, I went to Semargl.me, a Y-DNA database and searched using my Kit number (or my Ysearch number) and found him listed there at a Genetic Distance of 24).
Why Would I Take a BigY Test?
Family Tree DNA’s infographic about the BigY test, located above, provides some good reasons why someone would take a BigY test. It will potentially help contribute to our understanding of the human Y-DNA tree.
Additionally, there is incredible promise for members of family groups trying to tease apart relationships within the past 500, for example. These family groups are combining Y-STRs and Y-SNPs and BigY results to make some really interesting discoveries. But they are working with BigY results in a careful, planned manner.
If you are just starting out with Y-DNA and are on the fence about BigY, I would caution you to wait. This is a powerful tool for advanced users, but I don’t recommend that you purchase the test unless you have a more specific reason to do so. Instead, go out and test the autosomal DNA of 4 relatives!
What Else Can I do with My BigY Test?
The Family Tree DNA interpretation of your BigY results is not necessarily the end of the analysis! Once you have access to your BigY raw data, you can purchase an analysis of that raw data at YFull for $49 (see more about YFull on the ISOGG Wiki here).
Among other analyses, YFull provides:
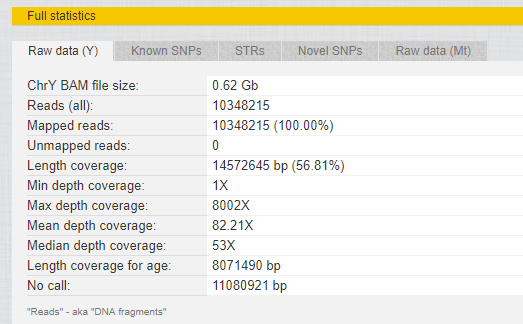
- Statistics about my raw data (see the screenshot below);
- Terminal SNP and location on the human Y-DNA tree (updated regularly);
- A list of Y-SNP genetic matches estimated to share a MRCA within the past 2,500 years;
- A list of novel SNPs found only in the test-taker;
- Results for 500+ Y-STRs (see the screenshot below); and
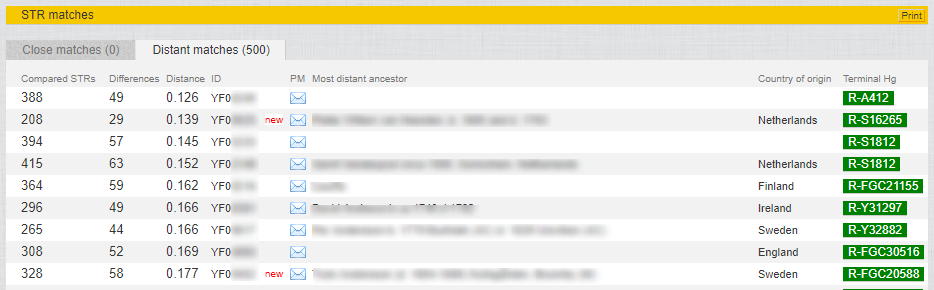
- A list of Y-STR matches (my closest is a GD=49 at 388 markers, and he has the same terminal SNP designation as I do).
My haplogroup and terminal SNPs:
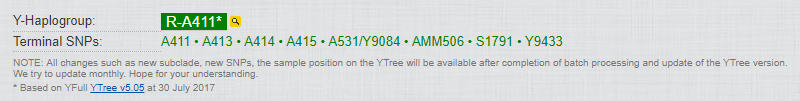
My statistics:
Some of my STR results:
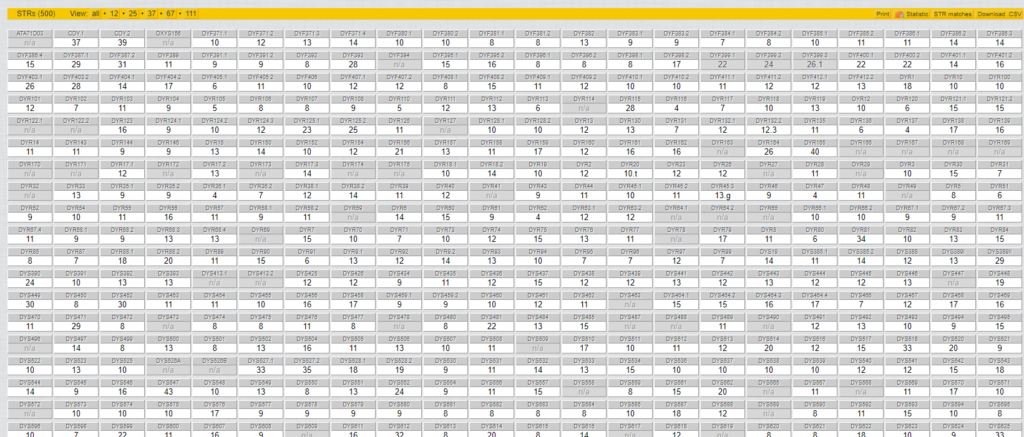
My SNP matches:
Some of my STR matches:
So what say you? Are you considering a BigY test? What do you hope to accomplish with your test?
Hi Blaine –
You’ve done it again. Thank you very much for a clear definition of BigY results and rationales for testing. Very timely as well. Would it be fair to say that BigY would help if you had many, many Y-DNA matches and wished to narrow it down? And conversely, when my Y-accounts for several people don’t even show many matches (other than 12 markers, which I tend to ignore), would it probably be unhelpful and therefore wise to wait?
If you have many Y matches at higher marker levels a Big Y would most certainly help. If you look at your Y matching table at your highest tested marker level do you see any common themes in your matches SNPs column? This could give you a clue on who has Big Y tested in your match list. I wish that the Y Matching table included a check mark or something for those that have Big Y tested. Anyways, if many of your matches are in a project you could email the project admin and ask for advice on whether or not a Big Y would be currently “worth it” to help delineate
your branching with your matches further.
I am administering a Y-DNA surname project. So far, members from 8 different families have tested with the 37-marker STR test. 7 of the 8 families are within a genetic distance of 2, with the MRCA going back to the early 1700’s. I am trying to get a better picture of the branching pattern within this tree within the last 300 years. Would you recommend BigY testing, 67-marker STR testing, targeted SNP testing, or some other plan of attack?
Thank you.
It can help. If you did not want to test all 8 members at $395 a pop you could have 1-2 members test. If they all happen to share and create new SNPs that were novel to themselves, which would likely be the case if they are suspected to share MRCA in 1700s, then you can take the furthest down (or most recent) SNP that they discovered in their Big Ys and SNP test the other 6 members to automatically place them down on this most recent SNP. If money is no issue there could possibly be some advantages to testing all 8 with a Big Y as some may end up having new novel SNPs that mutated after their shared common ancestor, but on average Big Y is able to resolve a SNP for about every 150 years. Some more expensive but higher resolution tests such as Full Genomes Corp Y Elite 2.1 are able to discover SNPs at about one per every 75 years. These differences are in the sequencing techniques, as the technology improves and becomes more affordable we will someday see new SNPs in the troublesome regions with high repeats (some of the same areas we test STRs). But since Big Y reads 100 base pair lengths at a time then goes back and rebuilds the whole strand against the reference genome, it fails to indentify SNPs in these regions. FGC Y Elite does 150 base pair reads, and new long read tech like FGC Chromium ($2950) reads in 50,000 or so base pair lengths so when reassembled there is less area and those crazy repeat regions can be resolved.
I took the test a couple years back. Ended up with my own SNP as a result. And one match with 6 Shared Novel Variants and 0 Known SNP Difference. And we share the same new SNP.
What is the estimated time difference between us with a GD of 10 at 111 markers?
still so expensive , i hope @FamilyTreeDNA let down its price
to 300 $ is more convinient for us ( in africain countries )
or make a new special offer to our countries in africa
Is the BigY test different from the Y-DNA 111 test at FTDNA?
Yes, the various markers (12, 25, 37, 67, 111) tests are single tandem repeats or sections of the Y chromosome where two to thirteen nucleotides repeated hundreds of times in a row on the DNA strand, e.g., “gatagatagatagata” would be a STR marker value of ‘4’ at a certain common location on the Y chromosome.
SNPs or single nucleotide polymorphisms are where a single nucleotide at a known location along the Y switches to another nucleotide. For example, L21, a large clade under R1b haplogroup, is a SNP at location ChrY: 15654428 (think of the Y strand as a interstate and this is a mile marker all our personal highways are compared to) where the base switched from the ancestral C to a G. Now all men descended from this ancestor that had this mutation some 4400 years ago have this SNP with the nucleotide or base at this location being a G.
SNP packs test already known locations for these mutations, Big Y test the majority of all these locations or all these mile markers along the Y (certain exceptions I won’t get into) and even looks at ones that have not been known to mutate in any testers to see if has happened to do so in your line. If it does find areas of your Y that are reliable and unique to you but a base changed, then someone else tests Big Y and shares this same change, it typically means you both descended from a common ancestor. Many of these new SNPs is how the haplotree is advanced with some family and surname projects even finding branching into genealogical relevant timeframes.
Quest Who Zack and Mike Kahn. Your all going to Jail for Hacking my DNA Tests and Hacking into my whole Life. SEE THE GREEN LINES UNDER THAT SENTENCE IT MEANS ITS TRUE WHAT I SAID. This is Kevin Lee Phillips , Aka Sir Edward Weldon Kahn, Kraus, Kaiser and all you hackers! Im coming for all of you with the FBI , CIA, Mr Trump . On my side because you people are Criminals.
Blaine, would you discuss competitive products, and pros/ cons of each? I’d hate to spend even this discount price, then find I really need to spend more hundreds of bucks on a different product. This is the first I’ve heard “since Big Y reads 100 base pair lengths at a time then goes back and rebuilds the whole strand against the reference genome, it fails to indentify SNPs in these regions.” (Thanks, Zack!)
Or can you give links to such discussions?
Thanks!
Hi Richard,
You can find information about competitive products here: https://isogg.org/wiki/Y-DNA_SNP_testing_chart
I had already done my y-DNA for 111 markers plus other analysis that showed that my y-DNA Haplogroup = R-BY3927.
I took advantage of the Big Y sale and just got back my results, which as I expected told me almost nothing. I predicted that none of my cousins (with common ancestors that lived within the past several centuries had bothered to do the test……since only 2 had done their y-DNA, and only one of those two had published his identity.
I figure that the payoff for doing my Big Y will (if ever) occur several decades after my demise.
This thread is great for the people across the globe and contains the informative stuff for the concerns.
Ordered Y-111. Got results. Closest matches were… genetic distance 1 on 25-marker. And it was from Italy. And, there were some GD2 matches on 25 marker, but… they all had either arabic or Nordic names. As if Arabs and Nordic were related…
They perdicted my haplogroup as R-M198. So I Ordered Big Y. Expecting the highest possible level of weirdness. Probably will turn out I’m from the Moon or Mars. Haplogroup Þ-Ö666 most likely.- :Ð
Hi Ms. Blaine! I could really use your professional advice in furthering my career and moving forward with a Master’s degree. I currently have hold a BS degree in Biology (with a simultaneous Pre-Vet medicine track and minor in Microbiology: I was two classes short of a second BS degree, ran out of money and time). I spent almost 2 years researching my mother’s side of the family which started in joining the DAR, and I couldn’t stop….I ended up getting her side back to the 4th Century AD. Like so many others, I fell in LOVE with geneology and found things I NEVER would have imagined (direct ancestry, grandparents line) to English and French royalty and the Holy Roman empire! As a Biologist I’ve had 2-3 courses in genetics…I can get financial aid for a Master’s degree but it has to be for a degreed program, AND I have to be able to work at least part-time as an older, single adult…guess you know the question coming ?,…what degrees Master’s program (and schools?) should I be looking for to be a genetic geneologist? I’m also a big fan of American and European history….Thanks SO much for any help you can provide…
awesome blog. good work admin
Hi, I ordered the Big y test several weeks ago and received conf and kit numbers by email but no sample vial yet in the post. Can you check for me when this will arrive? cheers rod Jeffery Kit St 111444
Hello, I am wanting to order two new Big Y DNA test for relatives. I have already had one cousin complete a test and have learnt that it is wise to have two more of mu cousins tested to provide extra information in order to bring the results closer to the 1600s. I am prepared to stand corrected on this and hope for clarification.
My questions are: 1; My cousin is the first tester and his son and grandson are both
prepared to participate in the tests.
2: Is this arrangement appropriate and how do I order two tests?? Maybe I just order two individually?
I am ordering and paying for these tests and manage their trees. Do I send them directly to the males involved???? Thank you for help and advice. Roslyn